A protein isolation method for western blot to study histones with an internal control protein
Abstract
Histone modification is one of the attractive targets for epigenetic studies. However, current methods to extract chromatin-associated proteins for western blot of histone modification have some weaknesses such as the loss of housekeeping proteins. In this study, we are presenting a simple method to isolate nuclear protein for studying histone modification by immunoblotting with housekeeping proteins. This method provided high protein concentration from minute tissue samples and importantly, it allowed us to detect acetylated histones together with internal control proteins such as β actin.
SHORT COMMUNICATION
Epigenetics has been emerging as an attractive field in biomedical sciences. Somatic changes within an individual or between individuals, which are not caused by changes in DNA sequence such as mutation, are promising targets for epigenetic studies, where DNA methylation and histone modifications are mostly considered. DNA methylation, one of the most important mechanisms controlling the gene expression, is established early in development [1]. Histone modifications regulating the manipulation and expression of DNA in most biological processes [2] which may occur throughout the life under different conditions.
To study the epigenetic regulation of gene expression, we have some good techniques such as western blot, DNA methylation assay, and chromatin immunoprecipitation (ChiP) assay. Western blot seems to be method considered as a simple and cheap technique to investigate global histone modifications applied in several epigenetic studies [3-4]. However, the traditional methods of nuclear protein isolation and histone extraction for immunoblotting of global histone modifications have some weaknesses including the low yield of isolated proteins and do not allow to have internal controls [3, 5]. Therefore, some additional steps and costs are needed to spend on preparing samples and analyzing the global histone modifications.
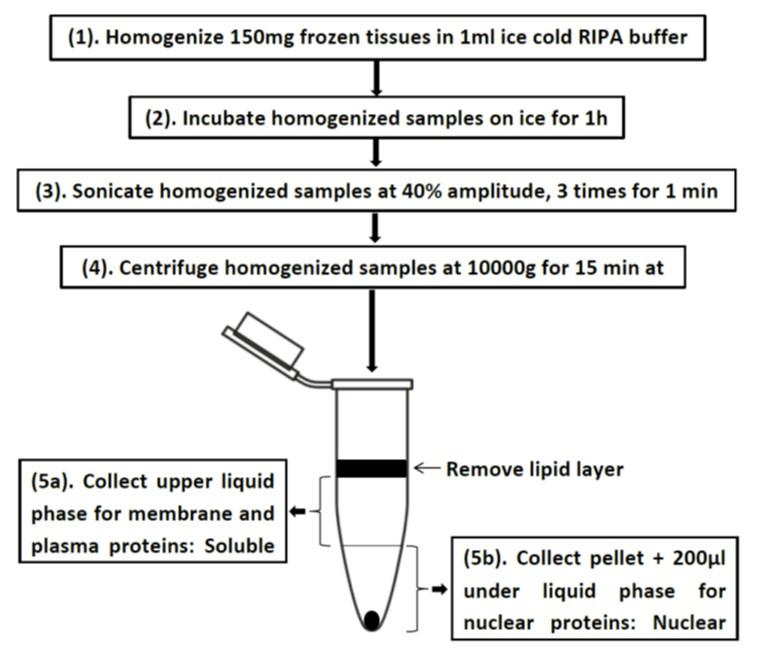
In this study, we have established a simple method (Fig. 1) to isolate nuclear proteins from lesser amounts of frozen tissues. This is a very simple method that produced not only the highest percentage of histones but also carried significantly high-level housekeeping proteins.
Concentrated protein isolated by our method and control method (acid extraction), was measured and compared by Bradford reagent as presented in Table 1. From the same amount of tissues (150mg/sample) and equal volume of dilution buffer (200ul/sample), our method yielded higher protein concentration than the acid extraction around 13 times or 29 times for inguinal fat (ING) or liver, respectively. But, the protein concentration was almost the same in the soluble (So) and nuclear (Nu) part of a homogenized sample isolated using our method. In our method, total proteins were collected with enriched histones in the nuclear part, whereas acid extraction tends to get pure nuclear proteins. However, in order to test the global histone modifications by western blot, the acid extraction method shows some weaknesses such as 1) it requires large amount of tissues for getting enough histones, 2) cannot run western blots with internal controls, and 3) isolated histones cannot be stored for a long period of time because they were extracted by 0.2N HCl. With such high protein concentration, our method showing low expressed acetylated histone 3. This result indicated that chromatins were abundant in the nucleus allowed us to run several western blots to check global modifications for all interested histones.
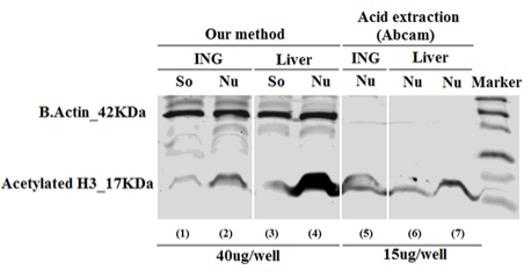
In order to confirm the enrichment of nuclear proteins are isolated by our method, we performed the western blot using a specific antibody against acetylated histone 3, has been considered as one of the most common chromatins. As shown in the Fig.2, the nuclear parts (Nu) of both inguinal fat (ING) (lane 2) and liver (lane 4) showing strong acetylated histone 3 expression, whereas the soluble parts (So) (lane 1 or 3) from same samples showing low expressed acetylated histone 3. This result indicated that chromatins were abundant in the nucleus allowed us to run several western blots to check global modifications for all interested histones.
Parts (Nu) isolated by our method, even we had similar protein concentrations in the Nu and So parts of the samples (Table 1). The expression of acetylatedhistone 3 was also found from the pure histones (lane 5, 6 and 7) purified by acid extraction method, because of low protein concentration was given by this method. We were unable to load more than 20µg protein/gel-well to get stronger signals, this is one of the weaknesses of acid extraction method.
Table 1. Protein concentration of different samples.
In the immunoblotting assay, it is better to have internal controls to confirm the equal amount of loaded proteins to normalize the quantified data. Internal controls (normalization controls) are the proteins of housekeeping genes such as β actin, glyceraldehyde-3-phosphate dehydrogenase (GAPDH), hypozanthine phosphoribosyltransferase 1 (HPRT1), and ribosomal protein large P1 (RPLP1) [11, 12]. Westernblots are used to study histones or their modifications use coomassie blue-stained gels to demonstrate equal loading [3, 5]. In our protein isolation method, we collected the part of samples having high chromatins, so our samples still had nuclear proteins as well as proteins of housekeeping genes. Therefore, this method allowed us to have the internal controls, such as β-actin in the blot (Fig. 2). As shown in Fig. 2, from lane (1) to (4); the signals of β-actin were strong and equal in both So and Nu parts of one tissue, and in both fat (ING) and liver samples. As expected, acid extraction methods did not give any signal of β-actin in both fat and liver samples (lane 5 to 7 of Fig. 2).
The results of the current study indicate that this alternative protein isolation method is a better choice to study chromatins and other proteins as well. Moreover, this method offers several advantages including small amount of tissue samples giving high protein concentration, one gel for both targeted and control proteins, and it was simple. However, this method is only suitable for studying global expressions of proteins by western blot not by other methods such as ChiP assay as it does not yield pure chromatins.
CONFLICTS OF INTEREST
The authors declare that they have no conflict of interest.
References
- [1]Patterson K, Molloy L, Qu W, Clark S. DNA Methylation: Bisulphite Modification and Analysis. J Vis Exp. 2011:e3170.
- [2]Bannister AJ, Kouzarides T. Regulation of chromatin by histone modifications. Cell Res. 2011;21:381-95.
- [3]Trojer P, Zhang J, Yonezawa M, Schmidt A, Zheng H, Jenuwein T, et al. Dynamic Histone H1 Isotype 4 Methylation and Demethylation by Histone Lysine Methyltransferase G9a/KMT1C and the Jumonji Domain-containing JMJD2/KDM4 Proteins. Journal of Biological Chemistry. 2009;284:8395-405.
- [4]Wellen KE, Hatzivassiliou G, Sachdeva UM, Bui TV, Cross JR, Thompson CB. ATP-Citrate Lyase Links Cellular Metabolism to Histone Acetylation. Science. 2009;324:1076-80.
- [5]Zhang Q, Ramlee MK, Brunmeir R, Villanueva CJ, Halperin D, Xu F. Dynamic and distinct histone modifications modulate the expression of key adipogenesis regulatory genes. Cell Cycle. 2012;11:4310-22.
- [6]Chu DT, Malinowska E, Gawronska-Kozak B, Kozak LP. Expression of Adipocyte Biomarkers in a Primary Cell Culture Models Reflects Preweaning Adipobiology. Journal of Biological Chemistry, 2014; 289(26): 18478-88.
- [7]Jura M, Jarosławska J, Chu DT, Kozak LP. Mest and Sfrp5 are biomarkers for healthy adipose tissue. Biochimie, 2016; 124: 124-33. [8] Chu DT, Malinowska E, Jura M, Kozak LP. C57BL/6J mice as a polygenic developmental model of diet-induced obesity. Physiological Reports, 2017;5(7): 20.
- [8]Chu DT, Malinowska E, Jura M, Kozak LP. C57BL/6J mice as a polygenic developmental model of diet-induced obesity. Physiological Reports, 2017;5(7): 20.
- [9]Rim JS, Mynatt RL, and Gawronska-Kozak B. Mesenchymal stem cells from the outer ear: a novel adult stem cell model system for the study of adipogenesis. The FASEB Journal, 2005.
- [10]Anunciado-Koza R, Ukropec J, Koza RA, Kozak LP. Inactivation of UCP1 and the glycerol phosphate cycle synergistically increases energy expenditure to resist diet-induced obesity. J Biol Chem, 2008. 283(41): 27688-97.
- [11]Aldridge GM, Podrebarac DM, Greenough WT, Weiler IJ. The use of total protein stains as loading controls: An alternative to high-abundance single-protein controls in semi-quantitative immunoblotting. Journal of Neuroscience Methods. 2008;172:250-4.
- [12]Gilda JE, Gomes AV. Stain-Free total protein staining is a superior loading control to β-actin for Western blots. Analytical Biochemistry. 2013;440:186-8.