Comparison of major nutritional constituents and genetic diversity analysis of five strains of oyster mushrooms
Abstract
Oyster mushroom is the second runner-up among commercially produced mushrooms due to its delicious taste, higher nutritional and medicinal properties. The objectives of this study were to determine the nutritional variation through the analysis of obtained nutritional values and to assess the genetic diversity through RAPD marker of five different strains of oyster mushrooms namely Pleurotus cystidiosus (strain: pcys2); Pleurotus djamor (pop1); Pleurotus ostreatus (ws); Pleurotus ostreatus (po3) and Pleurotus geesteranus (pg4). The strains showed variations in moisture, protein, fiber, lipid, ash and carbohydrate content ranged from 86.10-87.33%; 17.8-24.13 gm/100gm; 18.16-25.46 gm/100gm; 3.16-5.16 gm/100gm; 9.16-11.46 gm/100gm; and 35.4-45.33 gm/100gm respectively. In case of genetic diversity, the segregation of five strains of oyster mushrooms were grouped through un-weighted pair group method of arithmetic means average (UPGMA), where strains were grouped into two main clusters and the generated linkage distance was 48. The strains pop1 and po3 were aligned in cluster two (C2) due to their genetic similarity but showed dissimilarities with other strains. Though the strains pcys2, ws, and pg4 were aligned in the same cluster (C1), the strain ws was aligned in a different sub-cluster due to its few dissimilarities with the other two strains. The variation of nutritional values and genetic diversities among the mushroom strains indicates nutritional and genetic variabilities. The findings of current study indicate that, though these mushrooms were genetically dissimilar, all strains were nutritious with high protein and fiber contents with low fat. However, mushroom breeder can consider strain po3 for high protein content, strain ws for high fiber content and strain pop1 for low fat content.
INTRODUCTION
Mushroom is regarded as an ingredient of gourmet cuisine all over the world, particularly for its unique flavor and culinary wonder appreciated by humankind. Over 10,000 mushroom species exist in nature and around 200 mushroom species have served worldwide as functional foods for many years [1] while only 35 species have been cultivated commercially [2]. In recent decades, mushroom consumption has increased considerably due to scientific evidence that they help to fight and prevent a variety of diseases and helps the body to meet the nutritional requirements [3]. They are good source of non-starchy carbohydrates, high content of dietary fiber, moderate quantities of proteins and vitamins like ascorbic acid, niacin, riboflavin, and thiamine [4]; [5]. Mushrooms generate 20-35% of the protein dry weight, which is low in fats and contains all nine essential amino acids [6]. A number of secondary metabolites are accumulated by mushrooms including phenolic substance, polyketides, terpene, diboviquinone and steroids [7]. Additionally, mushrooms contain a large variety of bioactive compounds and have been shown to be especially efficacious as an antioxidant, antifungal, anti-cancer, anti-tumor, immuno-stimulant, and antimicrobial agents, which is seeking great attention from food and pharmaceutical sectors [8], [3], [9].
Pleurotus species require a short growth time compared to other mushrooms and are one of the highest-produced mushrooms in the world [10]. The genus Pleurotus is distributed throughout the world with 200 saprophytic species based on their temperature and tropical climate [11]. It is an edible mushroom that contains essential bio-active molecules, nutritional ingredients like essential proteins, carbohydrates, vitamins, calcium, and iron and also has numerous biological effects [12], [13]. Its extract can reduce cholesterol in the same way as dietary supplements [14]. Due to their extraordinary ligninolytic properties, Pleurotus (also known as oyster mushroom) is one of the most widely studied White-Rot fungi [8]. Pleurotus cultivation has the potency to be a highly profitable agribusiness because of its significant nutritional and therapeutic value [15].
DNA fingerprinting has emerged as a critical resource in the characterization of mushrooms [16]. The rapid generation of reliably reproducible DNA fragments is possible by using RAPD markers to a wide variety of mushroom species. Ravash et al. (2009) stated that the genomic wide and random nature of the RAPD technique is best to specify overall genetic affinity than the morphological study [17]. Alam et al., (2010) reported that RAPD able to generate polymorphism successfully that helps to discriminate of closely connected genotypes [18]. The proper identification of the Pleurotus sp. is important to leverage their full potential in the food industry [19]. Selecting high yield strains is vital for efficient cultivation. But, due to consecutive sub-culturing and production, the efficiency of commercial mushroom strains continue to decrease [20]. As toxic elements may present, a few more information on oyster mushrooms is required to detect proper strains for commercial production. Hence, the objective of this study was to analyze the nutritional values and to determine the genetic diversity of five strains of oyster mushrooms. The findings would help mushroom breeders in designing and developing good quality strains.
MATERIALS AND METHODS
Collection of strains
Fruiting bodies of five different species of oyster mushrooms namely P. cystidiosus (pcys2), P. djamor (pop1), P. ostreatus (ws), P. ostreatus (po3), and P. geesteranus (pg4) were obtained from culture house of the National Mushroom Development and Extension Centre, Savar, Dhaka, Bangladesh where the standard growing condition was ensured properly. The five species of fresh mushrooms were collected in clean polyethylene and were sun-dried. Then the dried samples were ground to fine powder by using a blender and finally with mortar and pestle, preserved in a desiccator until analysis. These dried samples were further dried in an oven at 60°C until a constant weight was obtained for elemental analysis. Samples were triplicated for each experiment and the experiment was repeated three times to reduce experimental errors.
Nutritional quality analysis of oyster mushrooms
Determination of moisture content
For the determination of moisture content, 5g of fresh mushroom was taken into an automatic moisture analyzer (Weighed moisture box, A & D company ltd. N 92; P1011656; Japan). The moisture content of five mushroom strains was determined according to Raghuramulu et al. (2003) [21]. In short, fresh mushrooms were dried at 100-105°C in an oven and were cooled in a desiccator. The heating and cooling were performed repeatedly until getting moisture-free mushroom. The following formula was used to calculate the moisture content percentages:
Moisture content (%) = (Initial weight-final weight) × 100/weight of sample
Determination of protein content
In order to determine protein content, five grams of grinded mushroom were mixed with 50 ml (1.25N) NaOH and allowed to boil for 30 minutes. The solution was left for cooling until the temperature reached 25 0C and it was then centrifuged in a tabletop centrifuge machine at 1000 x g. The total protein content of the collected supernatant was measured according to the Biuret method [22]. Briefly, the peptide bonds inside the proteins react with Cu2+ ions in alkaline solutions. Here the absorbance was measured by spectrophotometer (at 540 nm) and it was directly proportional to protein content.
Determination of lipid content
Lipid content was determined according to the method described by Folch et al. (1957) with a few modifications [23]. Briefly, five grams of grinded mushroom was suspended and mixed thoroughly in 50 ml of chloroform-methanol mixture (2:1 v/v) and the solution was kept for 3 days. The solution was then filtrated and centrifuged at 1000 x g. The methanol was removed from the upper layer by Pasteur pipette while chloroform was evaporated by heating. The rest substance was crude lipid, which was determined by the following formula:
Lipid content (g/100g sample) = Weight of lipid × 100/Weight of sample taken
Determination of fiber content
In the case of fiber content determination, five grams of moisture and the fat-free sample was taken into 200 ml of boiling H2S04 (0.255N) in a beaker and was mixed properly. The mixture was then boiled at a constant volume for 30 minutes by adding water continually. Therefore, the mixture was filtered through a muslin cloth, and the residue was washed with hot water until it was free from acid. The filtered material was transferred into the same beaker, adding 200 ml of boiling NaOH (0.313N). This boiling and washing step was repeated once again. The material was washed again with some alcohol and ether until it was free from alkali. It was then moved to a crucible and let dry overnight at 80-100° C and weighed (W1) in an electrical balance. The crucible was heated for 8 hours in a muffle furnace at 600° C, cooled, and weighed again (W2). The difference between initial and final weight (W1-W2) represents the weight of crude fiber.
Crude Fiber Content (g/100g sample) = [100 − (moisture + fat)]×(W1-W2)/Weight of sample [19].
Determination of ash content
Ash content was determined through the traditionally used method. Briefly, one gram of the sample was taken in a crucible and was placed on a triangle of clay pipes. It was then heated over a low flame until all the material had been fully charred, followed by heating around 8 hours in a muffle furnace at 600° C temperature. It was then kept in a desiccator for cooling and measuring. The crucible was heated again in the muffle furnace for 1 hour, cooled and measured, to ensure the completion of ash. The procedure was repeated until two consecutive weights were the same. Total ash content was calculated by the following formula:
Ash content (g/100g sample) = Weight of the ash × 100/ Weight of sample taken [19]
Determination of carbohydrate content
The content of the available carbohydrate was determined by the following formula:
Carbohydrate content (g/100g sample) = 100-[(Moisture + protein + Lipid + Fiber + Ash) g/100gm] [19]
Molecular characterization of mushroom
DNA extraction
Filamentous fungi have solid walls of the cells, which are tempting to rupture. The modified method of Aljanabi et al. (1999) was used to extract the genomic DNA from the mushroom [24]. DNA of five strains of oyster mushrooms was extracted from the 0.2-0.3 g fruiting body of each species. It was grinded with a mortar pestle in extraction buffer (200 mM Tris-HCl, pH-8.5, 250 mM NaCl, 0.5% SDS, 25 mM EDTA). The lysates were incubated for 40 minutes at 65°C in a water bath then centrifuged for 30 minutes at 10,000 x g. After adding an equal amount of isopropanol, DNA was precipitated and the resulting pellet was washed with 70% ethanol. The DNA pellet was air-dried and dissolved in 50μl TE buffer for long time storage at -20°C.
RAPD analysis
The Random Amplified Polymorphic DNA (RAPD) technique was carried out to determine the genetic diversity of mushroom samples. Genomic DNA was amplified by the RAPD method described by Williams et al. (1990) [25], where six different types of unspecified oligonucleotide primers (Operon Technologies Inc.) were used to generate amplified fragments (Table 1). RAPD method was used in the study because of available resources and also in the light of existent pieces of literature [26,27]. Besides, a recent study conducted by Familoni and colleagues used RAPD to determine the genetic diversity of wild type Pleurotus species in Nigeria [28]. The reaction mixture for each PCR reaction has consisted of 7.5 µl PCR master mix (containing Taq DNA polymerase, dNTPs, MgCl2, and reaction buffers), 1.5 µl primer, 2 µl DNA template, 4 µl nuclease-free water in a final reaction volume of 15 µl. PCR amplification was done in an oil-free thermal cycler (Piko Tm PCR machine), following the thermal cycle of 95°C for 2 minutes (initial denaturation) followed by 40 cycles of 30 sec denaturation at 95°C, 30 sec annealing at 37°C, and elongation or extension at 72°C for 1 minute where the final extension was 5 minutes at 72 °C. Finally, the PCR product was held at 4°C. At gel electrophoresis, 100bp and 1 kb DNA ladder were used to measure the DNA bands.
Table 1. The name of the primers and their sequences are listed below.
RAPD data scoring
Following electrophoresis, the size of amplification products was estimated by comparing the migration of each amplified fragment with that of a known size fragment of molecular marker l00bp to 1kb DNA ladder. To analyze the presence or absence of particular alleles, each amplification product produced by each RAPD primer was scored as ‘1’ or ‘0’, respectively. A cluster analysis UPGMA based on Nei’s and Li’s [29] was carried out using the ‘STATISTICA’ software [30]. The similarity and genetic distance between different species were estimated while a dendrogram was generated also.
Statistical analysis
Mean, standard deviation and ‘p’ value were calculated using standard statistical methods [31, 32].
RESULTS
Nutritional quality analysis in different oyster mushrooms
The present study was done to determine and compare nutrient contents of five strains of Oyster mushrooms such as P. cystidiosus (pcys2); P. djamor (pop1); P. ostreatus (ws); P. ostreatus (po3); P. geesteranus (pg4). The nutritional composition of different oyster mushrooms, including moisture, protein, lipid, fiber, ash, and carbohydrate was determined, shown in Table 2.
Table 2. Comparison of nutrient content among five strains of Oyster mushrooms.
Moisture content
The moisture content of the five strains namely P. cystidiosus (pcys2); P. djamor (pop1); P. ostreatus (ws); P. ostreatus (po3); P. geesteranus (pg4) were 86.53%, 87.33%, 87.06%, 87.23%, 86.10% respectively. There was no significant difference (P ˃ 0.05) among the values of moisture content of five strains of mushrooms.
Protein content
As mentioned in the Table 2, the protein contents (g/100g) determined in this study ranged from 17.76g to 24.13g. There was a significant difference (P ˂ 0.05) among the protein content of the strains.
Lipid content
In this study, the lipid content (g/100g) of P. cystidiosus (pcys2); P. djamor (pop1); P. ostreatus (ws); P. ostreatus (po3); P. geesteranus (pg4) mushrooms were found 5.16g, 3.16g, 3.66g, 4.10g, and 4.10g, respectively. Among all strains, P. cystidiosus (pcys2) contains the highest lipid mass (5.16g). Overall, Lipid content was found statistically significant (P ˂ 0.05).
Fiber content
In this research, an appreciable amount of fiber was present in the assessed mushrooms. The quantity of five different strains of oyster mushrooms (g/100g) fell within the range of 18.16g to 25.46g, which were also statistically significant (P ˂ 0.05).
Ash content
The Ash content of foods represents their mineral element composition. Mushrooms are strong bio accumulators, which is revealed by their medicinal attributes. The ash content of five different mushrooms was found 11.33g, 10.66g, 11.46g, 9.66g, 9.16g, respectively and statistically proven as significant (P ˂ 0.05).
Carbohydrate content
In this study, the carbohydrate content of five different mushrooms was found 42.16g, 45.33g, 35.36g, 39.33g, and 39.56g, for P. cystidiosus (pcys2); P. djamor (pop1); P. ostreatus (ws); P. ostreatus (po3) and P. geesteranus (pg4) respectively.
DNA fingerprinting using RAPD markers
DNA fingerprinting of five different mushroom strains was performed using six RAPD markers (OPA-03, OPF-01, OPA-09, OPAK-04, OPG-05, OPG-04) developed by Operon Tech., Inc., Alameda, California, USA. Among the six RAPD primers tested, OPA-03, OPAK-04, and OPG- 05 primers provided a relatively maximum amount of low smearing and high-intensity bands.
All the six RAPD primers used a total of 173 amplified bands from the five different oyster mushrooms using the Thermal Cycler (Piko Tm PCR machine). Images of electrophoresis using the primers OPA-03, OPF-01, OPA-09, OPAK-04, OPG-05, and OPG-04 were shown in Figure 1A, 1B, and 1C respectively. The primer, OPAK-04 amplified the highest number of 33 bands; primer, OPA-03 amplified a total of 32 bands; primer, OPF-01 amplified a total of 29 bands; primer, OPA-09 amplified a total of 28 bands; primer, OPG-05 amplified a total of 32 bands and primer, OPG-04 amplified the lowest number of 19 bands (Table 3). Out of the 173 bands, 151 bands were polymorphic, 3 bands were monomorphic bands and 19 bands were unique bands. The primers OPA-03, OPF-04, OPA-09, OPAK-04, OPG-05 and OPG-04 produced 30 (93.75%), 26(89.65%), 23(82.14%), 29(87.87%), 28(87.50%) and 15 (78.28%) polymorphic bands respectively (Table 3).
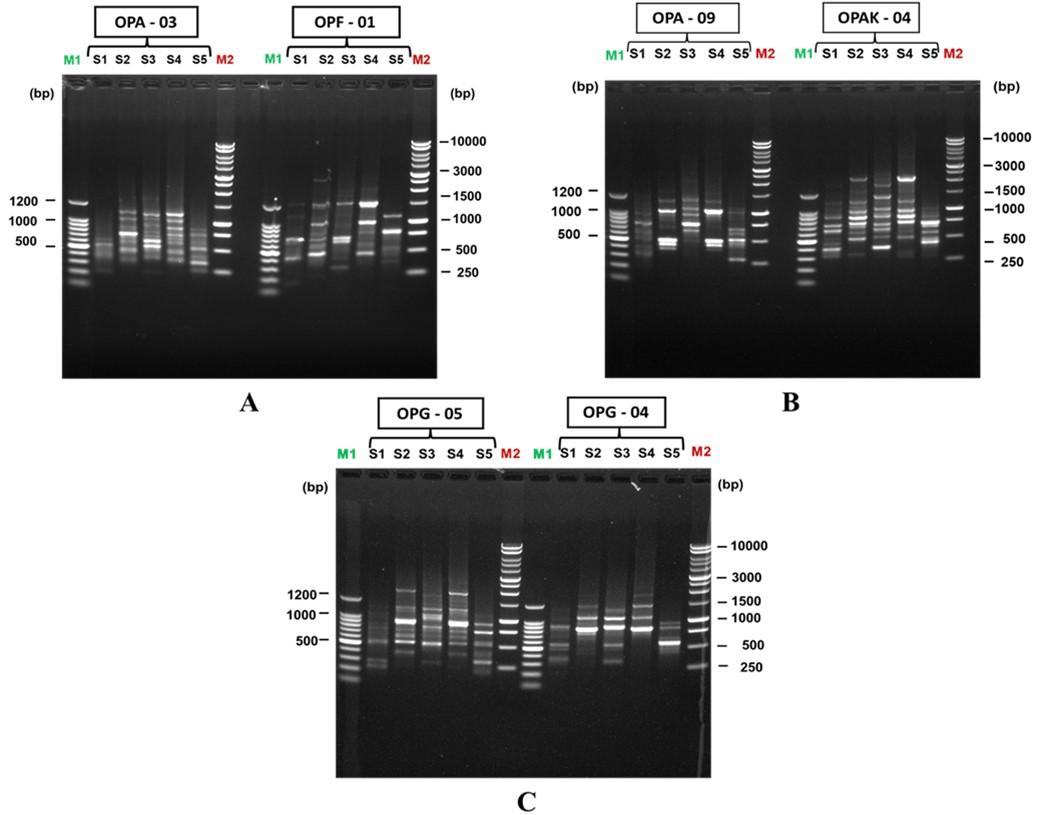
Table 3. RAPD primers with their corresponding DNA bands score in five different strains of oyster mushrooms.
Linkage distances
The values of pair-wise comparisons of the genetic distances scrutinized using the software program ‘STATISTICA’. The linkage distance among the strains was calculated from the cumulative data for the six primers ranging from 9 to 53. The highest linkage distance 53 was recorded in two cases between two strain pairs P. djamor (pop1) and P. ostreatus (ws). The lowest linkage distance 9 was recorded between the strains pairs P. djamor (pop1) and P. ostreatus (po3). Details of linkage distance were shown in Table 4.
Table 4. Summary of linkage distances for different pairs of five different oyster mushrooms using RAPD markers.
Cluster analysis (Tree diagram)
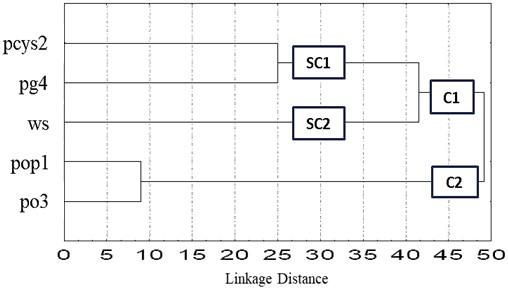
The RAPD based dendrogram revealed two major clusters by using the UPGMA. UPGMA indicated the segregation of five strains of oyster mushrooms into two main clusters (C1 and C2) presented in Figure 2 were produced at the linkage distance 48. P. djamor (pop1) and P. ostreatus (po3) in cluster two (C2) showed genetic similarity but were quite different from other strains. Further, the Dendrogram demonstrated that cluster one (C1) divided into two sub-cluster, where sub-cluster (SC2) P. ostreatus (ws) was different from sub-cluster (SC1) containing the strains P. cystidiosus (pcys2) and P. geesteranus (pg4).
DISCUSSION
Presence of different nutrient components in a standard level is a prerequisite of a healthy food. Hence, the nutrient content of five strains of oyster mushroom was observed and compared. The strains showed variations in moisture, protein, fiber, lipid, ash and carbohydrate content. The moisture content was ranged from 86.10-87.33% where the protein content ranged from 17.8-24.13 gm/100gm, fiber content ranged from 18.16-25.46 gm/100gm, lipid content ranged from 3.16-5.16 gm/100gm, ash content ranged from 9.16-11.46 gm/100gm and carbohydrate content ranged from 35.4-45.33 gm/100gm. The highest moisture, protein, lipid, fiber, ash and carbohydrate content were observed in the strains P. djamor (pop1), P. ostreatus (po3), P. cystidiosus (pcys2), P. ostreatus (ws), P. ostreatus (ws), and P. djamor (pop1) respectively while the lowest content were observed in the strains P. geesteranus (pg4), P. cystidiosus (pcys2), P. djamor(pop1), P. djamor (pop1), P. geesteranus (pg4), and P. ostreatus (ws) respectively.
High moisture contents promote susceptibility to microbial growth and enzyme activity [33].The moisture content percentage of oyster mushrooms was determined by Moni et al. (2004) [34] and Alam et al. (2008) [35] where the percentage were 88.15-91.64, and 87-87.5 respectively. Kumela and Solomon (2017) also found the moisture content 88.75% in 100g fresh mushrooms. These findings were quite similar to present study [36]. In case of protein content, in two different studies, Li et al. (2017) and Tolera and Abera (2017) reported the ranges of the protein of P. ostreatus mushroom was from 20.4g to 28.8g per 100g dried fruit bodies [12], [36], whereas Kortei et al. (2014) reported protein content ranging from 10.48% to 10.78% [33]. In another study, Ahmed et al. (2013) found lipid content ranged 3.5-4.7% in different oyster mushrooms [37]. The findings of other researches were also relevant to our findings [35,38]. Nevertheless, lower values were reported by some researchers as well [33,39,40], [41,42]. In case of fiber content , Ahmed et al. (2013) and Hoa et al. (2014) found (20.05-29.75)% dietary fiber in different oyster mushrooms, which were quite similar to our present study [37]; [41]. However, lower values are obtained by Patil et al. (2010) [39]. In case of ash content, Li et al. (2017), Patil et al. (2010), and Kortei et al. (2014) obtained the range of ash content from 5.90% to 9.9%, which is less than the range of our findings [12] [33,39]. In case of carbohydrate content, Dundar et al. (2008) and Hoa et al. (2015) evaluated the carbohydrate quantity of varied Pleurotus species ranged from 30.78 to 51.93 (g/100g), which were relevant to our findings [43], [41]. However, higher values were stated by several researchers as well [33,38,40].
Molecular markers, such as RFLP, RAPD, and genotyping have been used to discriminate mushroom species or strains of Agaricus, Auricularia, Ganoderma, Lentinula, Stropharia, Rugoso-annulata, and Volvariella. All of these technologies provided data for mushroom strain identification and protection [44]. Genetic diversity of mushrooms has been determined previously using molecular markers especially RAPD [28], [27], [45] and this technique is used to assess the genetic diversity among 37 Pleurotus species of mushrooms and found that it provides better discrimination than morphological analysis [46]. Besides, several published research used primers ranging from 3 to 8 to conclude their study [26,47], [48]. In this study, six RAPD primers were used and the highest linkage distance 53 was observed twice between the strain-pair P. djamor (pop1) and P. ostreatus (ws) while the lowest linkage distance 9 was found between the strain-pair P. djamor (pop1) and P. ostreatus (po3). The dissimilarity of the number of bands may be due to the primer’s sequence and availability of annealing sites in the genome [49] (Kernodle et al., 1993). Further, the Dendrogram demonstrated that cluster one (C1) divided into two sub-cluster, where sub-cluster (SC2) P. ostreatus (ws) was different from sub-cluster (SC1) containing the strains P. cystidiosus (pcys2) and P. geesteranus (pg4). Khan et al. (2011) reported that the formation of sub-groups in cluster A, by P-56 (Pleurotus sajor-caju) and P – 17 (Pleurotus florida), in cluster B, P-19 (Pleurotus ostreatus) and P-7 (Pleurotus flabelatus) and in cluster C, P-9 (Pleurotus warm-starm) and P-16 (Pleurotus .eryngii) are because of genetic distance 0.86, 0.80 and 0.81 respectively. They concluded that, the species within a distinct subgroup might be due to their same genus Pleurotus and same ancestry [50].
CONCLUSION
The current study suggests that the strains of oyster mushrooms differ from each other considering their nutritional composition although they are of the same genus, each species are nutritious with high protein and fiber content with low fat. The statistical analysis revealed that the protein, lipid, fiber, ash, and carbohydrate content of five strains of oyster mushroom were highly significant except for moisture content. In particular, the low lipid and high fiber contents of the oyster mushrooms make them health beneficial food items, especially against heart diseases and diabetes. The use of oyster mushrooms as food or ingredient for processed food products are promising because of their nutritional attributes and potential benefits for health. The six RAPD primers used in this study were able to identify the genetic diversity and to determine the phylogenetic relationship among the strains of Pleurotus species of oyster mushroom. This study further confirms that RAPD-PCR is a suitable tool for phylogenetic relationship determination and maintenance of good quality spawn of oyster mushrooms. All the strains used in this study were nutritious with high protein and fiber contents with low fat even though they were genetically dissimilar which were indicated through RAPD analysis. Hence, mushroom breeder can consider strain P. ostreatus (po3) for high protein content, strain P. ostreatus (ws) for high fibre content and strain P. djamor (pop1) for low fat content. In the future, this research work may help to determine the genetic relationship among other useful edible mushrooms as well.
ACKNOWLEDGEMENT
This work acknowledges the Research Laboratory, National Mushroom Development and Extension Centre, Savar, Dhaka, Bangladesh.
AUTHOR CONTRIBUTION
AS, DKP, NCS, and AJK contributed to study the design and did the study. AS, AR, and TE, collected the data. AS, SKB, SBS, and DKP contributed to data analysis and interpretation. AS, SKB, SBS, and AR drafted the article with the help of all authors. DKP, SST, SKB, TE, and NCS review the manuscript critically. DKP and NCS supervised the project. All authors read and approved the final manuscript.
CONFLICTS OF INTEREST
There is no conflict of interest among the authors.
References
- [1]Kalač P. A review of chemical composition and nutritional value of wild-growing and cultivated mushrooms. J Sci Food Agric 2013;93:209–18. https://doi.org/10.1002/jsfa.5960.
- [2]Aida FMNA, Shuhaimi M, Yazid M, Maaruf AG. Mushroom as a potential source of prebiotics: a review. Trends Food Sci Technol 2009;20:567–75. https://doi.org/10.1016/j.tifs.2009.07.007.
- [3]Heleno SA, Ferreira RC, Antonio AL, Queiroz MJRP, Barros L, Ferreira ICFR. Nutritional value, bioactive compounds and antioxidant properties of three edible mushrooms from Poland. Food Biosci 2015;11:48–55. https://doi.org/10.1016/j.fbio.2015.04.006.
- [4]Khatun S, Islam A, Guler P, Cakilcioglu U, Chatterjee NC. Hypoglycemic activity of a dietary mushroom Pleurotus florida (Mont.) Singer on alloxan induced diabetic rats. Biol Divers Conserv 2013.
- [5]Adebayo EA, Oloke JK, Azeez MA, Omomowo IO, Bora TC. Assessment of the genetic diversity among ten genotypes of Pleurotus (oyster mushroom) using nutrient and mineral compositions. Sci Hortic (Amsterdam) 2014;166:59–64. https://doi.org/10.1016/j.scienta.2013.12.010.
- [6]Kalač P. Chemical composition and nutritional value of European species of wild growing mushrooms: A review. Food Chem 2009;113:9–16. https://doi.org/10.1016/j.foodchem.2008.07.077.
- [7]Khatun S, Islam A, Cakilcioglu U, Guler P, Chandra N. NJAS – Wageningen Journal of Life Sciences Nutritional qualities and antioxidant activity of three edible oyster mushrooms ( Pleurotus spp .). NJAS – Wageningen J Life Sci 2015;72–73:1–5. https://doi.org/10.1016/j.njas.2012.03.003.
- [8]Bellettini MB, Fiorda FA, Maieves HA, Teixeira GL, Ávila S, Hornung PS, et al. Factors affecting mushroom Pleurotus spp. Saudi J Biol Sci 2019;26:633–46. https://doi.org/10.1016/j.sjbs.2016.12.005.
- [9]Khatun S, Islam A, Cakilcioglu U. Research on Mushroom as a Potential Source of Nutraceuticals : A Review on Indian Perspective 2012;2:47–73.
- [10]Park B, Ha BS, Kim MK, Lee B, Choi JI, Ryu J. Characterization of simple sequence repeats in the Pleurotus ostreatus cultivars , ‘ Heuktari ’ and ‘ Miso ’ 2016:174–8.
- [11]Carrasco-González JA, Serna-Saldívar SO, Gutiérrez-Uribe JA. Nutritional composition and nutraceutical properties of the Pleurotus fruiting bodies: Potencial use as food ingredient. J Food Compos Anal 2017;58:69–81. https://doi.org/10.1016/j.jfca.2017.01.016.
- [12]Li H, Zhang Z, Li M, Li X, Sun Z. Yield, size, nutritional value, and antioxidant activity of oyster mushrooms grown on perilla stalks. Saudi J Biol Sci 2017;24:347–54. https://doi.org/10.1016/j.sjbs.2015.10.001.
- [13]Yang Z, Xu J, Fu Q, Fu X, Shu T, Bi Y, et al. Antitumor activity of a polysaccharide from Pleurotus eryngii on mice bearing renal cancer. Carbohydr Polym 2013;95:615–20. https://doi.org/10.1016/j.carbpol.2013.03.024.
- [14]Khatun K, Mahtab H, Khanam PA, Sayeed MA, Khan AKA. Oyster mushroom reduced blood glucose and cholesterol in diabetic subjects: a short term intervention trial. Mymensingh Med J 2007;16.
- [15]Khatun K, Mahtab H, Khanam PA, Sayeed MA, Khan AKA. Oyster mushroom reduced blood glucose and cholesterol in diabetic subjects: a short term intervention trial. Mymensingh Med J 2007;16.
- [16]Shinwari ZK, Terauchi R, Kawano S. Molecular Systematics of Liliaceae — Asparagoideae — Polygonatae. I. RFLP Analysis of cpDNA in Several Asiatic Disporum Species. Plant Species Biol 1994;9:11–8. https://doi.org/10.1111/j.1442-1984.1994.tb00076.x.
- [17]Ravash R, Shiran B, Alavi A, Zarvagis J. Evaluation of genetic diversity in oyster mushroom (Pleurotus eryngii) isolates using RAPD marker. Journal of Science and Technology of Agriculture and Natural Resources. 2009;13(47 (B)):729-39.
- [18]Alam N, Lee J, Lee T. Mycelial growth conditions and molecular phylogenetic relationships of Pleurotus ostreatus. World Applied Sciences Journal. 2010;9(8):928-37.
- [19]Pawlik A, Janusz G, Koszerny J, Małek W, Rogalski J. Genetic diversity of the edible mushroom Pleurotus sp. by amplified fragment length polymorphism. Curr Microbiol 2012;65:438–45. https://doi.org/10.1007/s00284-012-0175-7.
- [20]Naraian R, Srivastava J, Garg SK. Influence of dairy spent wash (DSW) on different cultivation phases and yield response of two Pleurotus mushrooms. Ann Microbiol 2011;61:853–62. https://doi.org/10.1007/s13213-011-0206-9.
- [21]Raghuramulu N, Madhavan NK, Kalyanasundaram S. A Manual of Laboratory Techniques. Natl Inst Nutr Indian Counc Med Res Hyderabad-500 007, India 2003:56–8.
- [22]Burtis CA, Ashwood ER. Teitz Fundamentals of Clinical Chemistry (5th ed). Reed Elsevier India Priv Limited, New Delhi, India 2006:348–9.
- [23]Folch J, Lees M, Sloane Stanley G. A simple method for the isolation and purification of total lipids from animal tissues. J Biol Chem 1957;55:999–1033.
- [24]Aljanabi SM, Forget L, Dookun A. An Improved and Rapid Protocol for the Isolation of Polysaccharide- and Polyphenol-Free Sugarcane DNA. Plant Mol Biol Report 1999;17:1–8. https://doi.org/10.1023/A:1007692929505.
- [25]Williams JGK, Kubelik AR, Livak KJ, Rafalski JA, Tingey S V. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res 1990;18:6531–5. https://doi.org/10.1093/nar/18.22.6531.
- [26]Dwivedi S, Singh S, Chauhan UK, Tiwari MK. Inter and intraspecific genetic diversity (RAPD) among three most frequent species of macrofungi (Ganoderma lucidum, Leucoagricus sp. and Lentinus sp.) of Tropical forest of Central India. J Genet Eng Biotechnol 2018. https://doi.org/10.1016/j.jgeb.2017.11.008.
- [27]Alam N, Shim MJ, Lee MW, Shin PG, Yoo YB, Lee TS. Vegetative Growth and Phylogenetic Relationship of Commercially Cultivated Strains of Pleurotus eryngii based on ITS sequence and RAPD . Mycobiology 2009;37:258. https://doi.org/10.4489/myco.2009.37.4.258.
- [28]Familoni TV, Ogidi CO, Akinyele BJ, Onifade AK. Genetic diversity, microbiological study and composition of soil associated with wild Pleurotus ostreatus from different locations in Ondo and Ekiti States, Nigeria. Chem Biol Technol Agric 2018. https://doi.org/10.1186/s40538-018-0119-y.
- [29]Nei M, Li WH. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci U S A 1979;76:5269–73. https://doi.org/10.1073/pnas.76.10.5269.
- [30]Demidova L, Ivkina M, Zhdankina E, Krylova O, Sofyin E, Reshetova V, et al. Software Package STATISTICA and Educational Process. SHS Web Conf 2016;29:02011. https://doi.org/10.1051/shsconf/20162902011.
- [31]Lee DK, In J, Lee S. Standard deviation and standard error of the mean. Korean journal of anesthesiology. 2015 Jun;68(3):220.
- [32]McHugh ML. The chi-square test of independence. Biochemia medica. 2013 Jun 15;23(2):143-9.
- [33]Kortei NK, Dzogbefia VP, Obodai M. Assessing the Effect of Composting Cassava Peel Based Substrates on the Yield, Nutritional Quality, and Physical Characteristics of Pleurotus ostreatus (Jacq. ex Fr.) Kummer . Biotechnol Res Int 2014;2014:1–9. https://doi.org/10.1155/2014/571520.
- [34]Moni KH, Ramabardan R, Eswaran A. Studies on some physiological, cultural and post harvest aspects of Oyster mushroom Pleurotus ostreatus (Berk). Trop Agric Res 2004.
- [35]Alam N, Amin R, Khan A, Ara I, Shim MJ, Lee MW, et al. Nutritional Analysis of Cultivated Mushrooms in Bangladesh – Pleurotus ostreatus , Pleurotus sajor-caju , Pleurotus florida and Calocybe indica . Mycobiology 2008;36:228. https://doi.org/10.4489/myco.2008.36.4.228.
- [36]Tolera KD, Abera S. Nutritional quality of Oyster Mushroom (Pleurotus Ostreatus) as affected by osmotic pretreatments and drying methods. Food Sci Nutr 2017;5:989–96. https://doi.org/10.1002/fsn3.484.
- [37]Ahmed M, Abdullah N, Ahmed KU, Borhannuddin Bhuyan MHM. Yield and nutritional composition of oyster mushroom strains newly introduced in Bangladesh. Pesqui Agropecu Bras 2013;48:197–202. https://doi.org/10.1590/S0100-204X2013000200010.
- [38]Phan CW, Wong WL, David P, Naidu M, Sabaratnam V. Pleurotus giganteus (Berk.) Karunarathna & K.D. Hyde: Nutritional value and in vitro neurite outgrowth activity in rat pheochromocytoma cells. BMC Complement Altern Med 2012;12:1. https://doi.org/10.1186/1472-6882-12-102.
- [39]Sopanrao PATIL S, Abrar AHMED S, Manoharrao TELANG S, Mushtaq Vaseem BAIG M. the Nutritional Value of Pleurotus Ostreatus (Jacq.:Fr.) Kumm Cultivated on Different Lignocellulosic Agro-Wastes. Innov Rom Food Biotechnol 2010;7:66–76.
- [40]Akyüz M, Kirbaǧ S. Yenen yabani ve kültür mantarlarin besin deǧerleri. Turkish J Biol 2010;34:97–102. https://doi.org/10.3906/biy-0805-17.
- [41]Hoa HT, Wang CL, Wang CH. The effects of different substrates on the growth, yield, and nutritional composition of two oyster mushrooms (Pleurotus ostreatus and Pleurotus cystidiosus). Mycobiology 2015;43:423–34. https://doi.org/10.5941/MYCO.2015.43.4.423.
- [42]Abou Fayssal S, Alsanad MA, El Sebaaly Z, Ismail AIH, Sassine YN. Valorization of Olive Pruning Residues through Bioconversion into Edible Mushroom Pleurotus ostreatus (Jacq. Ex Fr.) P. Kumm. (1871) of Improved Nutritional Value. Scientifica (Cairo) 2020;2020. https://doi.org/10.1155/2020/3950357.
- [43]Dundar A, Acay H, Yildiz A. Yield performances and nutritional contents of three oyster mushroom species cultivated on wheat stalk. African J Biotechnol 2008;7:3497–501. https://doi.org/10.5897/AJB08.594.
- [44]Chandra S, Ghosh K, Acharya K. Comparative studies on the Indian cultivated Pleurotus species by RAPD fingerprinting. Nat Sci 2010;8.
- [45]Staniaszek M, Marczewski W, Szudyga K, Maszkiewicz J, Czaplicki A, Qian G. Genetic relationship between Polish and Chinese strains of the mushroom Agaricus bisporus (Lange) Sing., determined by the RAPD method. J Appl Genet 2002.
- [46]Stajić M, Sikorski J, Wasser SP, Nevo E. Genetic similarity and taxonomic relationships within the genus Pleurotus (higher Basidiomycetes) determined by RAPD analysis. Mycotaxon 2005.
- [47]Irfan M, Yang S, Yuxin L, Sun JX. Genetic diversity analysis of Morchella sp.by RAPD. Mol Biol Res Commun 2017;6:27–31. https://doi.org/10.22099/mbrc.2017.3955.
- [48]Ahlawat OP, Gupta P, Kamal S, Dhar BL. Variability in intra-specific and monosporous isolates of Volvariella volvacea based on enzyme activity, ITS and RAPD. Indian J Microbiol 2010. https://doi.org/10.1007/s12088-010-0031-z.
- [49]Kernodle SP, CANNON E, Scandalios JG. Rapid and simple phage DNA isolation. Bio techniques. 1993;14(3):360-2.
- [50]Khan SM, Nawaz A, Malik W, Javed N, Yasmin T, ur Rehman M, Qayyum A, Iqbal Q, Ahmad T, Khan AA. Morphological and molecular characterization of Oyster mushroom (Pleurotus spp.). African Journal of Biotechnology. 2011;10(14):2638-43.