Morphology, diversity and phylogenetic analysis of Spodoptera exigua (Lepidoptera: Noctuidae) in North Sulawesi by employing partial mitochondrial cytochrome oxidase 1 gene sequences
Abstract
Spodoptera exigua (Hübner, 1808) (Lepidoptera: Noctuidae) is a significant agricultural crop pest in Indonesia, causing significant economic losses in recent years. This species’ ability to survive on a wide variety of host plants provides an adaptive advantage for survival in the environment, which is facilitated by its high mobility, fecundity, and capability to acquire resistance to a broad spectrum of chemical pesticides. It is well-established that knowledge of diversity and evolutionary origins facilitate the development of pest management strategies. In the present study, we report the morphology, diversity and phylogeny analysis of S. exigua from North Sulawesi, Indonesia. The specimen from Rurukan have a body size and other segments that are longer than in Langowan and Modoinding. Dendrogram analysis shows that the similarity distance based on morphology ranges from 1-25%, which forms four clusters, where the specimen from Rurukan is separated from the rest of the specimens. The phylogeny of S. exigua from North Sulawesi, Indonesia, based on CO1 (Cytochrome c oxidase subunit 1) gene fragment, which is juxtaposed with CO1 data of the allied species from many geographical locations. A total of twenty-five isolates representing Indonesia, Japan, Germany, Thailand, India, UK, USA, Spain and Australia were compared. Nineteen sequences of S. exigua retrieved from GenBank were selected as references based on previous published phylogenic trees. The twenty-four isolates were scattered in two distinct clades indicating S. exigua is polyphyletic, but S. exigua from North Sulawesi, Indonesia is monophyletic.
INTRODUCTION
The beet armyworm, Spodoptera exigua Hübner (Lepidoptera: Noctuidae), has a wide range of distribution throughout the world, including Indonesia. It is considered a worldwide distribution pest due to its migratory capacity, which significantly contributes to the population outbreak and facilitates the geographic expansion of the population [1,2]. This species is native to Southeast Asia, has established itself as a significant insect pest of edible vegetables, and It is also resistant to a variety of insecticides [3]. The species is a polyphagous that may feed on more than 50 plant species belonging to more than ten plant families worldwide, including soybean, sugar beet, cabbage, cauliflower, brussels sprout, tomato, maize, cotton, lettuce, peanut, alfalfa, shallot, pastures crops, and various wild hosts [4].
In Indonesia, the pest was first reported attacking shallot plants in Java in December 1930 in large numbers. Currently, the pest has spread to virtually all parts of Indonesia, including Java, Sumatra, Sulawesi, Kalimantan, Bali, and East Nusa Tenggara. This insect has the capacity to spread rapidly on shallot plants in both the highlands and lowlands and is unaffected by the seasons throughout the year [5]. Based on the level of damage caused by this pest, it generally occurs in onion plants. Hence, it is called an important pest on onion plant [6,7]. Also, the insect is an obstacle to increasing crop yields of scallion due to a decrease in yield quantity and quality of 57-100% [8].
Study of the S. exigua in Indonesia has been conducted on population [6], pest management [9], life cycle [10], invasion and attack level [11], control using insecticides [9,12], and natural enemies [13]. The phylogeny of the Indonesian S. exigua, on the other hand, has not yet been reported in any publications. The knowledge of whether this insect was introduced into Indonesia and/or evolved locally implicates to practical pest management. Therefore, in the present study, we investigated the phylogeny of Indonesian S. exigua from North Sulawesi, Indonesia, and compared it with data on the same species in several geographical locations in the world. Evolutionary inferences were made by constructing gene genealogies from partial DNA sequences of the mitochondrial gene cytochrome c oxidase subunit 1 (COI).
MATERIALS AND METHODS
Sample collection
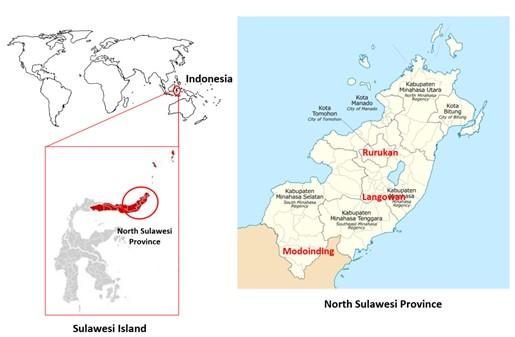
The specimen of S. exigua were collected during night time (07.00 – 08.00 p.m.) from different hosts, namely shallots, corn and peanuts in three different regions in North Sulawesi, Indonesia, including Tomohon City, Minahasa Regency and South Minahasa Regency (Figure 1). Adult insects (imago) were collected and stored in glass bottles, covered in gauze, and transported to the laboratory for morphological observation and molecular identification of the specimens.
Multivariate morphometric analysis
Observations of morphological characteristics were conducted using the approach described by Balvin et al. [14]. The qualitative parameters observed included eye color, wing color, wing pattern, body color, and body hair intensity. Quantitative parameters which are morphometric characters included head length (HL), thorax length (THL), abdomen length (ADL), antenna length (ANL), eye diameter (ED), wing length (WL), wingspan (WS), femur length (FL) and tibia length (TL). Observations of these morphological characteristics were carried out with a HIROX KH-8700 Digital Microscope (Hirox, Europe). Multivariate clustering analysis for morphometric characters was performed using SPSS IBM 20.
DNA extraction
Total DNA was extracted from fresh specimens using the innuPrep DNA Micro Kit (Analytik Jena) according to the manufacturer’s instructions. To achieve higher concentration of DNA yield, a slight modification was done according to Kolondam [15] by increasing the time for incubation in lysis solution (and proteinase K) to one hour. The purified DNA samples were preserved in the freezer (-20°C).
Polymerase chain reaction (PCR)
The PCR of samples were carried out using MyTaq HS Red Mix (Bioline) PCR kit. Every 40 µl reaction contained 20 µl of PCR premix, 15 pmol of each primer used, and 1 µl of DNA sample. Autoclaved MilliQ water was used to complete the volume to 40 µl. The primers employed in this research were based on Folmer et al. [16] as follows: LCO1490 (5′-GGT CAA CAA ATC ATA AAG ATA TTG G-3′) as forward primer and HC02198 (5′-TAA ACT TCA GGG TGA CCA AAA AAT CA-3′) as reverse primer. The Thermocycler (TPersonal, Biometra) setting was 95°C (3 minutes) for initial denaturation and continued with 35 cycles of 95°C (20 seconds) of denaturation, 50°C (30 seconds) of primer annealing, and 72°C (20 seconds) of DNA elongation. DNA separation of PCR products were done in agarose gel electrophoresis (0.8%) contained ethidium bromide. The PCR products were visualized using UV light to detect the 710 bp amplified band in PCR reaction.
DNA sequencing
The Sequencing were conducted by BigDyeTM Terminator v3.1 cycle sequencing kit chemistry in ABI PRISM 3730xl Genetic Analyzer (Applied Biosystems) by First Base C.O. (Malaysia). All of the samples were sequenced bi-directionally with both primers used in the PCR reaction. Chromatograms were assembled using MUSCLE algorithm [17], and edited under Geneious v5.6 [18] platform. The 658 bp long fragment generated were used for data analysis.
Molecular identification of specimens
Sequences of local S. exigua were deposited in GenBank. The GenBank accession numbers of Indonesian S. exigua as well accompanied data are shown in Table 1. Identification was accomplished through the use of the BLAST identity search feature supplied by the same platform (https://blast.ncbi.nlm.nih.gov/Blast.cgi).
Table 1. Sources and accession of S. exigua (No. 1-24;), and S. litura (No. 25) used in this study.
Phylogenetic analysis
The sequences of Indonesian S. exigua were aligned with reference sequences obtained from GenBank using the multiple alignment program CLUSTALW (v.1.83) plug-in integrated in Geneious v.5.3.6. The alignment was edited manually using Geneious v.5.3.6 and all polymorphisms were confirmed by re-examining the electropherograms. The evolutionary history was inferred by the Maximum Likelihood (ML) method based on the Kimura 2-parameter (K2P) model [19]. Evolutionary analyses were conducted in MEGA v10.0.4 [20].
RESULTS
Morphometric analysis of S. exigua from North Sulawesi
The morphometric study employed specimens from three localities in North Sulawesi, namely Tomohon, Minahasa, and South Minahasa. Visual examinations using a magnifying equipment revealed similarities in the morphology of moths from three districts in Tomohon City (Rurukan), Minahasa Regency (Langowan), and South Minahasa Regency (Modoinding). The color of the forewings was grayish brown, while the hind wings were white and slightly brownish, black eyes, antennae like threads and at rest the position of the wings like a precarious arrangement on the abdomen. However, as seen in Table 2, the average morphometry of all specimens varied.
Morphological characteristics that can be used in general as a marker or identification of this moth species are the wings. The forewings are rather narrow, while the hind wings are broad, and are mostly covered by feathers or scales, have irregular stripes and black spots on the sides. The outer edge of the wing is dark, with dark irregular stripes with yellow-orange spots. Some have silver spots on the forewings, quadruped cubital hind wings, and long and slender filamentous antennae in males and females, which varies in the color of the thorax and back. The color of the abdomen and the intensity of the hair on the organs, especially the legs, the appendages on the genitalia in the form of a collection of feathers lighter in color than the abdomen.
Table 2. Average morphometry (mm) of specimens from Rurukan, Langowan, and Modoinding.
Cluster analysis of S. exigua from North Sulawesi
Cluster analysis is a statistical clustering method used to analyze large amounts of data divided into clustered segments [21]. Clustering has been widely used since the 1990s aimed at grouping objects based on the similarity of characteristics between these objects. In this study, cluster analysis was carried out to determine the similarity of the morphological characteristics of S. exigua in different habitats, namely Rurukan, Langowan, and Modoinding. Each of these areas was represented by three samples with nine main morphological characters used, namely head length (HL), thorax length (THL), abdomen length (ADL), antenna length (ANL), eye diameter (ED), wing length (WL), wingspan (WS), femur length (FL) and tibia length (TL). The morphological character variables have distances which are arranged in Table 3.
The distance matrix table shows the distances between variables in nine morphological characters of S. exigua in North Sulawesi represented by three onion-producing centers, namely Tomohon (Rurukan), Minahasa (Langowan) and South Minahasa (Modoinding). The smaller the Euclidean distance, the more similar the two morphometric character variables were analyzed together from the three habitats of the S. exigua moth. The more similar characters will form a cluster (Table 4).
Following cluster analysis, groups are determined based on the degree of similarity. The dendrogram analysis of S. exigua insect samples from Rurukan, Langowan, and Modoinding using SPSS IBM ver. 21.0 resulted in a similarity index as shown in Figure 2.
The dendrogram study of insects from Rurukan, Langowan, and Modoinding revealed that the distance of similarity based on morphology was between 1 and 25%, forming four groups. The first cluster, consisting of insects from Langowan (L1, L2, L3) and Modoinding (M1, M2, M3), demonstrated morphological resemblance by creating a single group with a similarity level of 1%. The second cluster, insects from Rurukan (T1 and T2), also demonstrated morphological similarity at a 1% similarity index, resulting in the formation of a new group distinct from the first cluster. At a 2% similarity index, the third grouping, insects from Rurukan (T1 and T3), demonstrated morphological resemblance. Meanwhile, the fourth cluster demonstrates that the third cluster, S. exigua from Rurukan (T1 and T3), forms a single group with the first cluster, at a 25 percent similarity index.
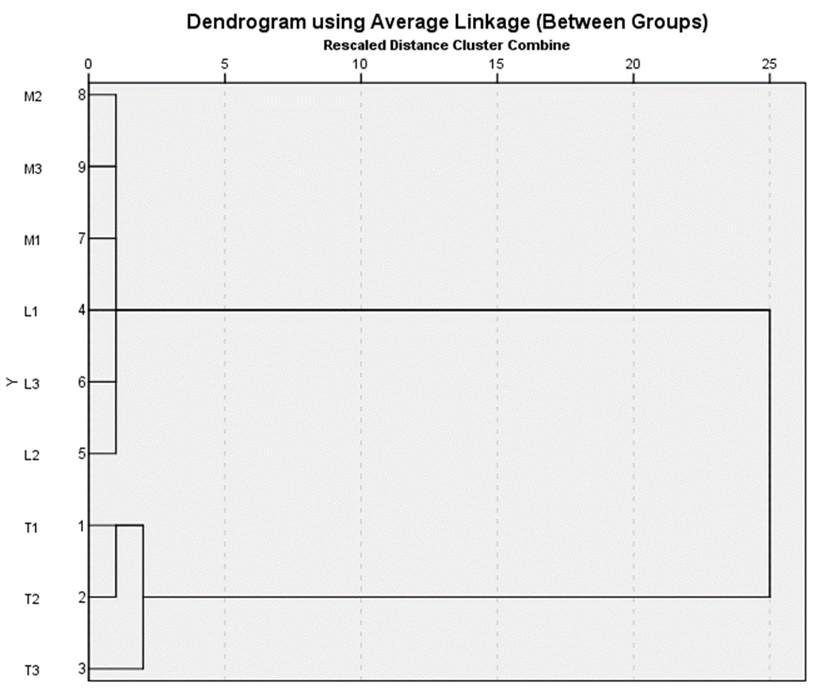
Table 3. Distance between variables of morphological character of S. exigua from Rurukan, Langowan and Modoinding.
Table 4. Formation of the morphological cluster of S. exigua from North Sulawesi.
Phylogeny analysis of S. exigua using the CO1 gene
This CO1 gene has been frequently utilized in insect research to assess inter- and intraspecific genetic variation [22]. It is also used to supplement standard morphological-based species identification for more precise findings [23,24]. Figure 3 depicts a molecular phylogenetic study using the neighbor joining method and the K2P model. The tree reveals that specimens form two clades, while specimens from North Sulawesi are clustered in one clade. Clade I include S. exigua from Japan (3 specimens), Germany (3 specimens), Thailand (1 specimen), Indonesia (6 specimens), India (1 specimen), UK (3 specimens), USA (3 specimens), and one specimen from Spain, with strong bootstrap support (100 %). Clade II is made up of only three specimens of the Australian S. exigua with bootstrap support of 100 %.
Table 5 summarizes the genetic distances between the specimens examined. S. exigua (HQ950504) from Australia and S. litura (HQ950413) from the Northern Territory of Australia have a genetic distance of 0.075. The farthest genetic distance between S. exigua was between specimens HQ950504 (Australia) and JF415658 (Germany), which was 0.052. Meanwhile, the genetic distance between S. litura (HQ950413) and S. exigua varies between 0.071 and 0.073.
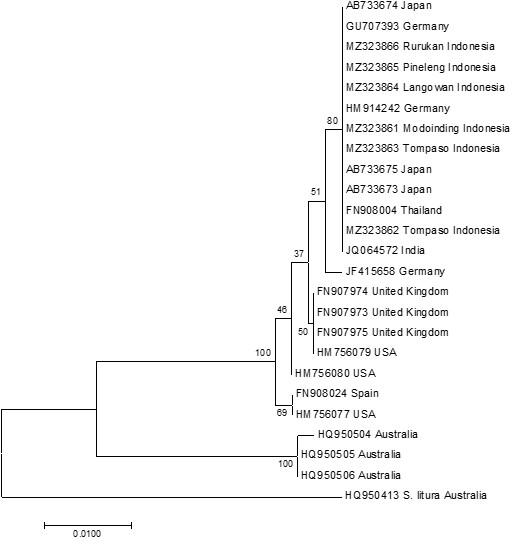
Table 5. Estimation of evolutionary divergence amongst S. exigua and S. litura specimens based on K2P method. The number of base substitutions per site from between sequences are shown in the lower diagonal; the number of base differences per site from between sequences are shown on the upper diagonal.
DISCUSSION
Despite extensive use of agrochemicals, insect pests continue to pose a significant threat to agricultural output and yields. For this reason, correct identification of the species is critical in the establishment of integrated pest control programs (IPM). Kinship relationship may be used to quantify the degree of similarity between species or populations [25]. This is in contrast to the diversity coefficient, which is used to quantify the degree of variation across species or populations on a set of characteristics [26]. It may be deduced from this connection that the further the kinship relationship, the greater the amount of variety and the lesser the level of uniformity, and vice versa.
In comparison to other animal species, insects have the greatest phenotypic plasticity within the animal group [27]. As a result, even within a single species, numerous morphological differences were discovered in insects. Environmental factors such as habitat and ecosystem conditions at the sampling location greatly impact the factors that determine the size difference across several populations [28]. Morphological variety in insects occurs as a result of environmental variables such as food and climatic availability, geographic position, and the existence of natural enemies, as well as the process of natural selection [29]. Morphological diversity occurs mostly in the shape and size of bodily organs in insects [30].
Another aspect that contributed to the Rurukan specimens having the lowest degree of kinship with the Langowan and Modoinding specimens was the considerably lower intensity of pesticide usage to suppress S. exigua on leek plants in the Rurukan region. This was feasible due to the low prevalence of S. exigua infestation on leek plants in the Langowan and Modoinding regions. According to farmers in Langowan and Modoinding, leeks were frequently treated with pesticides four to five times during the growing season, but farmers in Rurukan sprayed insecticides just two to three times during the growing season. According to Benítez et al. [29], morphological variation is widespread among insects because it is associated with adaptation to the environment in which they exist. S. exigua that resides in the Rurukan was assumed to undergo character changes, both in terms of nature and appearance or phenotype. These are referred to as environmental variants since they are generated only by changes in the environment while the genetic information stays unchanged. Environmental effects might manifest themselves in the form of climate variables and established agricultural practices [31].
While morphometric study revealed morphometry variation across S. exigua specimens from North Sulawesi, genetic distance analyses using the K2P method revealed no genetic variation among all of these specimens. This indicates that the environment has an effect on the morphological distinctions between the specimens without creating any intra-specific variation. Thus, S. exigua from North Sulawesi exhibited solely morphological plasticity.
Intra-specific genetic variation among specimens in Clade I ranges from 0.000-0.008, whereas among specimens from Australia (Clade II), it ranges from 0.000-0.002. According to Ashfaq et al. [32], intraspecific divergences in 81 butterfly species in North-central Pakistan ranged from 0.0 to 1.6% with a mean of 0.2%. Based on these findings, it may be concluded that the specimens of S. exigua examined in this study are most likely no longer S. exigua, since intraspecific variation has reached 0.052 (5.2%). As a result, a revisiting of the species S. exigua based on a more comprehensive morphometric investigation and multi-barcode regions is highly suggested.
Result of our analyses confirms that S. exigua seems well differentiated into two clades and that may correspond to two distinct species. The clade one constitutes a first putative species cluster whereas the clade two from Australia constitute another one. It is supported by Barcoding of life database (BOLD) that S. exigua specimens from Australia are grouped into a distinct barcode cluster (BOLD:AAA6645) that differ from another barcode cluster (BOLD:AAA6644) grouping all remaining S. exigua individuals. The result of this study is consistent with Shashank et al. [33] and Dumas et al. [34]. They reported that S. exigua is divided into two cluster groups and the Australian population of S. exigua is entirely different from other populations. This result requires more study using specific gene in order to determine whether these specimens correspond to a new species or to a case of cryptic species complex. The present study of phylogeny has confirmed that S. exigua is polyphyletic, whereas S. exigua from North Sulawesi, Indonesia is monophyletic.
CONCLUSIONS
The specimens from Rurukan have a body size and other segments that are longer than in Langowan and Modoinding. The similarity distance based on morphology ranges from 1-25%, which forms four clusters, where the specimen from Rurukan is separated from the rest of the specimens. Morphometric study revealed morphological variation across S. exigua specimens from North Sulawesi. However genetic distance analyses using the K2P method revealed no genetic variation among all of these specimens. The phylogeny of S. exigua from North Sulawesi, Indonesia, based on the CO1 (Cytochrome c oxidase subunit 1) gene fragment, which is juxtaposed with the CO1 data of the allied species from many geographical locations, is polyphyletic. However, S. exigua from North Sulawesi, Indonesia is monophyletic.
ACKNOWLEDGEMENT
The authors wish to thank the Ministry of Research, Technology and Higher Education for providing financial support for this research through the Doctoral Dissertation Research Scheme, Fiscal Year 2018.
AUTHORS CONTRIBUTION
Conceptualization, U.S., M.T. and A.P.; Methodology, U.S., J.P. and C.L.S.; Formal analysis, T.E.T. and A.P.; Data curation, U.S. and B.J.K.; Writing-original draft preparation, U.S., A.P., B.J.K., T.E.T. and T.B.E.; Writing- review and editing, A.P, T.E.T. and T.B.E.; Visualization, U.S. and T.E.T.; Supervision, M.T., J.P. and T.E.T.; Critical revisions and writing, T.E.T., A.P. and T.B.E. All authors have read and agreed to the published version of the manuscript.
CONFLICTS OF INTEREST
There is no conflict of interest among the authors.
References
- [1]Feng HQ, Wu KM, Cheng DF, Guo YY. Radar observations of the autumn migration of the beet armyworm Spodoptera exigua (Lepidoptera: Noctuidae) and other moths in northern China. Bull Entomol Res. 2003;93(2):115–24.
- [2]Fu X, Feng H, Liu Z, Wu K. Trans-regional migration of the beet armyworm, Spodoptera exigua (Lepidoptera: Noctuidae), in North-East Asia. PLoS One. 2017;12(8):e0183582-e0183582.
- [3]de Castro AA, Legaspi JC, Tavares W de S, Meagher RLJ, Miller N, Kanga L, et al. Lethal and behavioral effects of synthetic and organic insecticides on Spodoptera exigua and its predator Podisus maculiventris. PLoS One. 2018;13(11):e0206789.
- [4]Sarnthoy O. Comparative study of artificial diet and soybean leaves on growth, development and fecundity of beet armyworm, Spodoptera exigua (Hubner) (Lepidoptera : Noctuidae). Kasetsart Journal Nat Sci. 2000;34:339–44.
- [5]Moekasan T. Penerapan ambang pengendalian organisme pengganggu tumbuhan pada budidaya bawang merah dalam upaya mengurangi penggunaan pestisida. J Hortik. 2012;22(1):47.
- [6]Rauf A. Dinamika populasi Spodoptera exigua (Hubner) (Lepidoptera: Noctuidae) pada pertanaman bawang merah di dataran rendah. Bul Hama dan Penyakit Tumbuh. 1999;11(2):39–47.
- [7]Azidah AA. Population study of Spodoptera exigua (Lepidoptera: Noctuidae) larva and its affecting factors in Sekinchan, Selangor. Pakistan J Biol Sci PJBS. 2007;10(13):2152–8.
- [8]Putrasamedja S, Setiawati W, Lukman L, Hasyim A. Penampilan beberapa klon bawang merah dan hubungannya dengan intensitas serangan organisme pengganggu tumbuhan. J Hortik. 2016;22(4):349.
- [9]Aldini GM, Trisyono YA, Wijonarko A, Witjaksono W, De Putter H. Farmers’ practices in using insecticides to control Spodoptera exigua infesting shallot Allium cepa var. aggregatum in the shallot production centers of Java. J Perlindungan Tanam Indones. 2020;24(1):75.
- [10]Farahani S, Talebi AA, Fathipour Y. Life table of Spodoptera exigua (Lepidoptera: Noctuidae) on five soybean cultivars. Psyche (Stuttg). 2012;2012:513824.
- [11]Lestari AD, Suryanto A, Trisilowati. Dynamics of caterpillar pests (Spodoptera exigua) and purple blotch transmission in Lembah Palu red onion plants. AIP Conf Proc 2020;2264(1):50002.
- [12]Moekasan TK, Murtiningsih R. Pengaruh campuran insektisida terhadap ulat bawang Spodoptera exigua Hubn. J Hortik 2010;20(1):67–79.
- [13]Nusyirwan. Study of natural enemy Spodoptera exigua on onion agroecosystem. J Penelit Pertan Terap. 2013;13(1):33-37.
- [14]Balvín O, Munclinger P, Kratochvíl L, Vilímová J. Mitochondrial DNA and morphology show independent evolutionary histories of bedbug Cimex lectularius (Heteroptera: Cimicidae) on bats and humans. Parasitol Res. 2012 Jul;111(1):457–69.
- [15]Kolondam B. Applying matk gene for identification of liliopsida plant species fromnorth sulawesi through Bold systems. Int J Appl Biol Pharm. 2015;6(2):242-245
- [15]Kolondam B. Applying matk gene for identification of liliopsida plant species fromnorth sulawesi through Bold systems. Int J Appl Biol Pharm. 2015;6(2):242-245
- [16]Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotechnol. 1994;3(5):294–9.
- [16]Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotechnol. 1994;3(5):294–9.
- [17]Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32(5):1792–7.
- [17]Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32(5):1792–7.
- [18]Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics. 2012;28(12):1647-1649.
- [18]Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics. 2012;28(12):1647-1649.
- [19]Kimura M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol. 1980;16(2):111-120.
- [19]Kimura M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol. 1980;16(2):111-120.
- [20]Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol Biol Evol. 2018;35(6):1547–9.
- [20]Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol Biol Evol. 2018;35(6):1547–9.
- [21]Omran MGH, Engelbrecht AP, Salman A. An overview of clustering methods. Intell Data Anal. 2007;11(6):583–605.
- [21]Omran MGH, Engelbrecht AP, Salman A. An overview of clustering methods. Intell Data Anal. 2007;11(6):583–605.
- [22]Caterino MS, Cho S, Sperling FA. The current state of insect molecular systematics: a thriving Tower of Babel. Annu Rev Entomol. 2000;45:1–54.
- [22]Caterino MS, Cho S, Sperling FA. The current state of insect molecular systematics: a thriving Tower of Babel. Annu Rev Entomol. 2000;45:1–54.
- [23]Tallei T, Koneri R, Kolondam B. Sequence analysis of the cytochrome c oxidase subunit 1 gene of Pseudagrion pilidorsum (Odonata: Coenagrionidae). Makara J Sci. 2017;21:43–52.
- [23]Tallei T, Koneri R, Kolondam B. Sequence analysis of the cytochrome c oxidase subunit 1 gene of Pseudagrion pilidorsum (Odonata: Coenagrionidae). Makara J Sci. 2017;21:43–52.
- [24]Lombogia CA, Posangi J, Pollo HN, Tulung M, Tallei TE. Assessment of genetic variation in Apis nigrocincta (Hymenoptera: Apidae) in Sulawesi revealed by partial mitochondrial cytochrome oxidase 1 gene sequences. Scientifica. 2020;2020:1609473.
- [24]Lombogia CA, Posangi J, Pollo HN, Tulung M, Tallei TE. Assessment of genetic variation in Apis nigrocincta (Hymenoptera: Apidae) in Sulawesi revealed by partial mitochondrial cytochrome oxidase 1 gene sequences. Scientifica. 2020;2020:1609473.
- [25]Ochoa A, Storey JD. Estimating FST and kinship for arbitrary population structures. PLoS Genet. 2021;17(1):e1009241.
- [25]Ochoa A, Storey JD. Estimating FST and kinship for arbitrary population structures. PLoS Genet. 2021;17(1):e1009241.
- [26]Tamaki I, Setsuko S, Tomaru N. Genetic variation and differentiation in populations of a threatened tree, Magnolia stellata: factors influencing the level of within-population genetic variation. Heredity. 2008;100(4):415-423.
- [26]Tamaki I, Setsuko S, Tomaru N. Genetic variation and differentiation in populations of a threatened tree, Magnolia stellata: factors influencing the level of within-population genetic variation. Heredity. 2008;100(4):415-423.
- [27]Moczek AP. Phenotypic plasticity and diversity in insects. Philos Trans R Soc Lond B Biol Sci. 2010;365(1540):593–603.
- [27]Moczek AP. Phenotypic plasticity and diversity in insects. Philos Trans R Soc Lond B Biol Sci. 2010;365(1540):593–603.
- [28]Khaliq A, Javed M, Sohail M, Sagheer M. Environmental effects on insects and their population dynamics. J Entomol Zool Stud. 2014;2:1-7.
- [28]Khaliq A, Javed M, Sohail M, Sagheer M. Environmental effects on insects and their population dynamics. J Entomol Zool Stud. 2014;2:1-7.
- [29]Benítez HA, Püschel T, Lemic D, Čačija M, Kozina A, Bažok R. Ecomorphological variation of the wireworm cephalic capsule: studying the interaction of environment and geometric shape. PLoS One. 2014;9(7):e102059.
- [29]Benítez HA, Püschel T, Lemic D, Čačija M, Kozina A, Bažok R. Ecomorphological variation of the wireworm cephalic capsule: studying the interaction of environment and geometric shape. PLoS One. 2014;9(7):e102059.
- [30]Jiang L, Hua Y, Hu G-L, Hua B-Z. Habitat divergence shapes the morphological diversity of larval insects: insights from scorpionflies. Sci Rep. 2019;9(1):12708.
- [30]Jiang L, Hua Y, Hu G-L, Hua B-Z. Habitat divergence shapes the morphological diversity of larval insects: insights from scorpionflies. Sci Rep. 2019;9(1):12708.
- [31]Skendžić S, Zovko M, Živković IP, Lešić V, Lemić D. The impact of climate change on agricultural Insect pests. Insects. 2021;12(5).
- [31]Skendžić S, Zovko M, Živković IP, Lešić V, Lemić D. The impact of climate change on agricultural Insect pests. Insects. 2021;12(5).
- [32]Ashfaq M, Akhtar S, Khan AM, Adamowicz SJ, Hebert PDN. DNA barcode analysis of butterfly species from Pakistan points towards regional endemism. Mol Ecol Resour. 2013;13(5):832–43.
- [32]Ashfaq M, Akhtar S, Khan AM, Adamowicz SJ, Hebert PDN. DNA barcode analysis of butterfly species from Pakistan points towards regional endemism. Mol Ecol Resour. 2013;13(5):832–43.
- [33]Shashank PR, Thomas A, Ramamurthy V V. DNA barcoding and phylogenetic relationships of Spodoptera litura and S. exigua (Lepidoptera: Noctuidae). Florida Entomol. 2015;98(1):223–8.
- [33]Shashank PR, Thomas A, Ramamurthy V V. DNA barcoding and phylogenetic relationships of Spodoptera litura and S. exigua (Lepidoptera: Noctuidae). Florida Entomol. 2015;98(1):223–8.
- [34]Dumas P, Barbut J, Le Ru B, Silvain J-F, Clamens A-L, d’Alençon E, et al. Phylogenetic molecular species delimitations unravel potential new species in the pest genus Spodoptera Guenée, 1852 (Lepidoptera, Noctuidae). PLoS One. 2015;10(4):e0122407–e0122407.
- [34]Dumas P, Barbut J, Le Ru B, Silvain J-F, Clamens A-L, d’Alençon E, et al. Phylogenetic molecular species delimitations unravel potential new species in the pest genus Spodoptera Guenée, 1852 (Lepidoptera, Noctuidae). PLoS One. 2015;10(4):e0122407–e0122407.