Granzyme B gene polymorphisms and risk of hepatocellular carcinoma in patients with chronic hepatitis
Abstract
Infection with the hepatitis B virus (HBV) continues to be a hazard for public health across the globe. Chronic hepatitis, cirrhosis, and hepatocellular carcinoma (HCC) are all possible outcomes. It is obvious that certain patients with chronic hepatitis B (CHB) viral infection developed HCC, while other under almost similar circumstances do not. The present study aimed to investigate the possible a link between three single nucleotide gene polymorphisms (SNPs) in GzmB genes with the development of HCC. A total of 85 patients diagnosed with CHB participated in this research (40 patients with HCC and 45 patients without HCC). Three SNPs in GzmB gene (rs7144366, rs8192917 and rs2236338) were genotyped using restriction fragment length polymorphism (RFLP). The haplotype blocks derived from the three SNPs were assembled, and the linkage disequilibrium (LD) between the SNPs was determined using the SHEsis software. The homozygous mutant genotype (CC) was shown to be significantly more common in patients with HCC (27.5 %) than in those without HCC (11.11 %) (OR= 3.93, 95% CI=1.13-13.62, p=0.031). At allelic level, the mutant allele (C) was more frequent in patients with HCC than those without HCC (46.25% vs. 26.67%) with a significant deviation (OR=2.36, 95%CI= 1.25- 4.49, p= 0.008). The haplotype block CCG was more common among patients with HCC (26.25%) than those without HCC (12.22%) with a significant difference (OR= 2.56, 95%= 1.14-5.71, p= 0.022). The study indicated that individuals carrying the mutant homozygous (CC) of the SNP rs8192917 and allele C of this SNP may have a higher chance of developing HCC compared with those carrying other genotypes and T allele of the SNP. The haplotype block CCG (corresponding for C allele of rs7144366, C allele of rs8192917 and G allele of rs2236338) might be regarded as a risk factor for the emergence of HCC in patients with CHB.
INTRODUCTION
Approximately 90% of all initial liver malignancies are triggered by hepatocellular carcinoma (HCC), prevalent in the world’s most populated areas. It accounts for 9.2 % of all new cancer cases and the third biggest cause of cancer-related death, and it is the fifth most prevalent cancer in men and the seventh most common cancer in women in the world, following only stomach and lung cancers [1]. The activation of various cellular receptors (such as toll-like receptors (TLR)-3, TLR-9, retinoic acid-inducible gene (RIG)-1) results in the rapid production of antiviral cytokines such as interferon (IFN), which, in cooperation with natural killer (NK) cells, limits the initial spread of the hepatitis B virus (HBV) [2]. The humeral arm of immune response may also participate in body defense against the virus through the maturation and proliferation of B-cells to create antibodies specific to hepatitis B antigen [3]. The granules of cytotoxic T lymphocyte (CTL) and NK cells are discharged into the immunological synapse that has been established with their target cells. Serine protease granzymes (Gzms) are the granules’ most potent contents, which are transported into the cytoplasm of the target cells by the pore-forming protein perforin and trigger programmed cell death [4]. Granzyme B is the most important member of granzymes. It causes early DNA damage by causing the death of target cells through a mitochondria-dependent mechanism [5]. CTL responds to viruses infected cells by producing Gzms. Those Gzms should work at maximum speed with lower Km and should be free of the inhibitors for the enzymes to initiate apoptosis successfully [3]. The objective of the current investigation was to investigate the possibility of a relationship between three single nucleotide gene polymorphisms in the GzmB gene and the development of HCC in patients with CHB.
MATERIALS AND METHODS
Study population
This study was conducted from 1 January to 31 December 2021, in the Department of Chemistry and Biochemistry at Al-Nahrain University’s College of Medicine, in collaboration with the Department of Biochemistry Laboratory at Al-Imamain Alkadhimain Teaching Hospital and Medical Research Unit/College of Medicine/Al-Nahrain University. The Institutional Review Board (IRB) of the College of Medicine at Al-Nahrain University gave its clearance to the research project (number 202012158 on 19 January 2021). A total of 85 patients with CHB were included in the study. Those patients were split up into two different groups. Group 1 included 45 patients diagnosed with hepatocellular carcinoma, and group 2 included 45 patients without hepatocellular carcinoma.
The diagnosis of CHB was primarily based on positivity for HBsAg and anti-HBc-IgG antibodies accompanied by viral load beyond the reference range. The diagnosis of HCC depended on ultrasound (US) findings and Computed tomography (CT) scan. Patients with renal failure, connective tissue disease, acute and chronic infection, hepatitis C virus (HCV), and other malignancies were excluded from the study.
Blood samples
Venous blood (about 5 ml) was collected from all subjects, with 2 mL in an EDTA tube and 3 mL in a plan tube. The serum was extracted from the latter after centrifugation and stored at -20 ºC until used.
HBV serological markers
Ready commercial kits were used to detect HBsAg, anti-HBc IgG, HBeAg, and anti-HBe using the commercial ELISA technique (Bioelisa/biokit/Barcelona/Spain).
DNA extraction and gene amplification
Following the manufacturer’s instructions, a ready kit (gSYNCTM DNA Mini Kit Whole Blood Protocol/ Geneaid/ Korea) was used to extract DNA from blood samples. The GzmB gene fragments involving rs7144366, rs8192917, and rs2236338 gene polymorphisms were amplified using three sets of primers (Table 1).
The final concentration of 100 pmole/L was achieved by dissolving the primers in DNase and RNase free water, which was used in the experiments. Mixing 10 μL of the stock solution with 90 μL of distilled water resulted in a working solution with a concentration of 10 pmol/L. Each master mix tube included 4 μL of template DNA and 2 μL of primers. The amplification of GzmB gene segments matching to the three SNPs was carried out under the circumstances of a flowing PCR reaction. The annealing temperature was the sole variable; it was 58, 60, and 61 oC for the rs7144366, rs8192917, and rs2236338 gene polymorphisms, respectively. The first denaturation was performed at 95oC for 5 minutes, followed by denaturation at 94oC for 30 seconds, annealing for 30 seconds, elongation at 72 oC for 1 minute, and final elongation at 72 oC for 7 minutes.
Table 1. Primers and their corresponding single nucleotide polymorphisms.
Restriction fragment length polymorphism
The PCR product was digested with the restriction enzyme. The restriction enzymes and their cutting sequence are illustrated in Table 2.
Five μL of PCR products were mixed with 0.1μL of the selected restriction enzyme (Sibenzyme, Russia). Restriction buffer (10X) 1.5 µL and bovine serum albumin (BSA) 0.1µL were added to the mixture. The reaction mixture was then completed to 15µ l by molecular grad water. Mineral oil (20 µl) was added to each tube to prevent evaporation. The reaction mixture was incubated in 60◦ C water bath for 3 hours. Restriction products were resolved on 2% agarose.
Table 2. The restriction enzymes, their cutting sequence, and fragment length resulted from digestion.
PCR product sequencing
Ten percent of the PCR products were sent to the Macrogen Company in South Korea to verify RFLP for direct sequencing. The generated nucleotide sequences were put through a BLAST alignment process, which compared them to the reference sequence. This tool is accessible online at http://www.ncbi.nlm.nih.gov.
Statistical analysis
Statistical analysis was performed using SPSS 25.0 (SPSS, Chicago). The student t-test was used. The genotype divergence from Hardy-Weinberg Equilibrium (HWE) was assessed using Chi-square. The OR and 95% confidence interval (CI) were calculated using binary logistic regression (CI). SHEsis software was used to create haplotypes and calculate LD between the GzmB gene polymorphisms rs7144366, rs8192917, and rs2236338. A statistically significant difference was defined as a p-value less than 0.05.
RESULTS
Demographic and laboratory data of the study population
Although patients with HCC demonstrated higher mean age than those without HCC (42.7±14.18 years vs. 37.8±13.46 years), the difference was not significant. Both groups were similar in their characteristics in terms of gender distribution and HBeAg positivity, with no significant differences. However, patients without HCC had far more frequent positivity for hepatitis B antigen and antibody (HBeAb) than those with HCC (46.67% vs. 7.5%), with a highly significant difference (Table 3-5). Interestingly, all patients in both groups were positive for HBsAg and HBcAb-IgG.
Table 3. Characteristics of the participants, both demographic and laboratory-based.
Molecular assays
Three SNPs were investigated in the present study for their association with HCC. The genotyping was performed by RFLP. The different genotypes in all included SNPs were in good accordance with HWE in both groups (with and without HCC).
rs7144366
Gel electrophoresis of the PCR product of rs7144366 polymorphism is shown in Figure 1. The fragment length was 97bp. Digestion with SfcI restriction enzyme revealed three genotypes: TT, CT, and CC as shown in Figure 2 and 3.
Although the homozygous mutant genotype (CC) was more frequent in patients with HCC than those without HCC (27.5% vs. 13.33%), the difference was not significant. Likewise, a higher percentage of HCC patients had the allele C (45%) than those without HCC (35.56%), with no significant difference (Table 4).

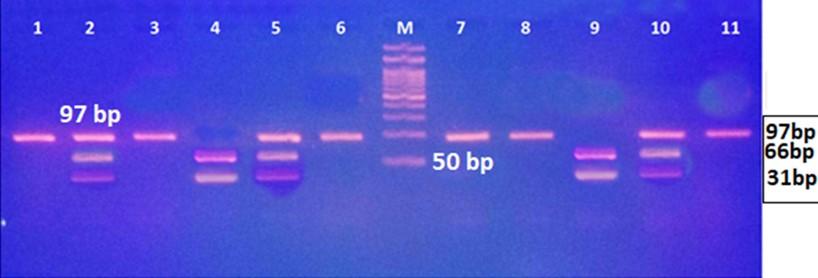
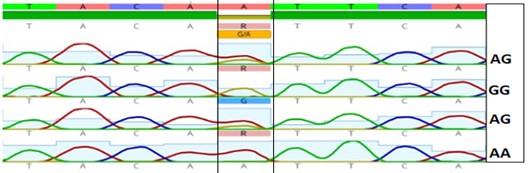
Table 4. The frequency of different genotypes and allele of rs7144366 in hepatitis patients with and without HCC.
rs8192917
Gel electrophoresis of the PCR product of rs8192917 polymorphism is shown in Figure 4. The fragment length was 420bp. Digestion with BsmA I restriction enzyme revealed three genotypes: TT, CT, and CC, as shown in Figure 5 and 6.
The homozygous mutant genotype (CC) was far more frequent in patients with HCC (27.5%) than those without HCC (11.11%) with a significant difference (OR= 3.93, 95%CI=1.13-13.62, p=0.031). This polymorphism seems to have recessive inheritance as the CC+CT genotypes were more common in patients with HCC than those without HCC (65% vs. 42.22%) with a significant difference (OR= 2.54, 95%CI= 1.06-6.12). At allelic level, the mutant allele (C) was more frequent in patients with than those without HCC (46.25% vs. 26.67%) with a notable distinction (OR=2.36, 95%CI= 1.25- 4.49, p= 0.008) as shown in Table 5.
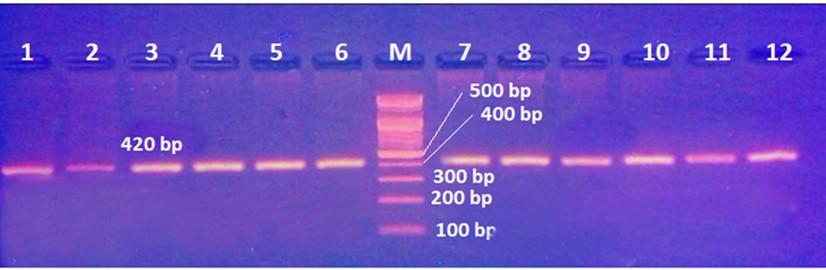
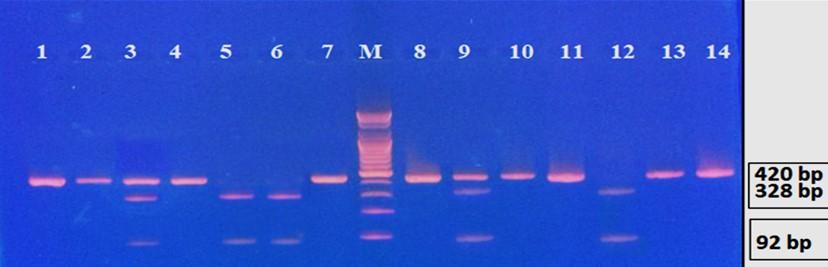
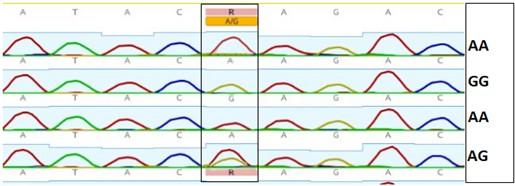
Table 5. The prevalence of various genotypes and alleles of rs8192917 in hepatitis patients with and without HCC.
rs2236338
Gel electrophoresis of the PCR product of rs2236338 polymorphism is shown in Figure 7. The fragment length was 400bp. Digestion with BsmF I restriction enzyme revealed three genotypes: AA, AG and GG as shown in Figure 8 and 9. There were no significant differences in the frequency of various genotypes and alleles of this polymorphism between individuals who had HCC and those who did not have the condition (Table 6).
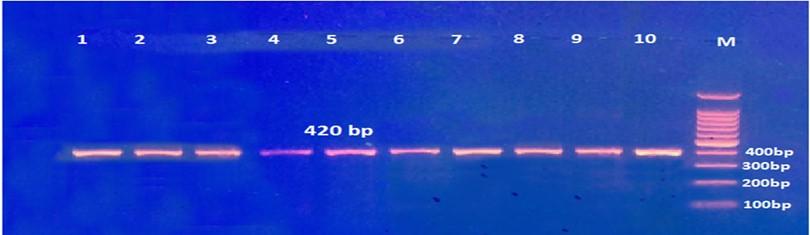
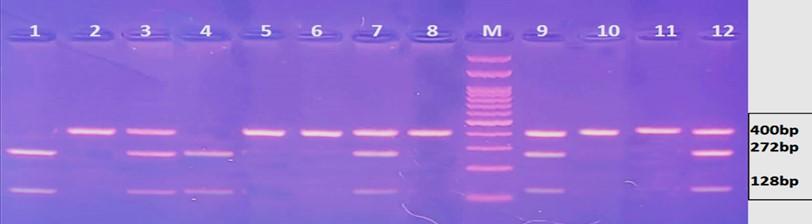
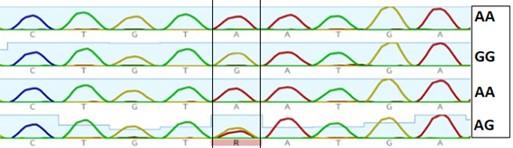
Table 6. The occurrence of various genotypes and alleles of rs2236338 in hepatitis patients with and without HCC.
Haplotype analysis
The SHEsis program was used to build haplotype blocks since the three polymorphisms are situated on the same gene. Table 7 shows the most frequent haplotypes in patients with and without HCC. The haplotype block TTA (T allele of rs7144366, T allele of rs8192917, and A allele of rs2236338) among patients without HCC had a higher prevalence than in HCC patients. (43.33% vs. 22.5%) with highly significant difference (OR= 0.38, 95%CI= 0.19-0.74, p= 0.005). in contrast, the haplotype block CCG was more common among patients with HCC (26.25%) than those without HCC (12.22%) with a significant difference (OR= 2.56, 95%= 1.14-5.71, p= 0.022). Although the haplotype block TTG was more common in patients with HCC (22.5%) than those without HCC (12.22%), the difference was not significant.
Table 7. The most frequent haplotype blocks.
Linkage disequilibrium
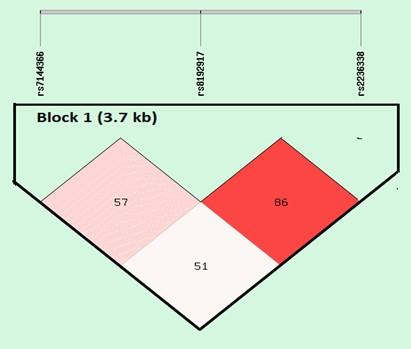
The results of LD analysis are presented in Figure 10. LD plot was constructed using combined genotype data from all patients. The SNP rs8192917 is in tight link with rs2236338 and rs8192917 (the measure D’ was 0.86). Furthermore, there was weak LD between rs8192917 and rs7144366 (the measure D’ was 0.57).
DISCUSSION
According to the result of the present study, the CC genotype of the SNP rs8192917 was significantly correlated with the development of HCC in patients with hepatitis B viral infection (OR= 3.93, 95%CI=1.13-13.62, p=0.031). This indicates that HBV patients with the CC genotype of this polymorphism are approximately four times more likely to develop HCC than individuals with the TT genotype. Furthermore, patients carrying the C allele of this polymorphism are at a 2.36-time higher risk than those with the T allele. On the other hand, the other two SNPs demonstrated no significant connection with the development of HCC in patients with HBV infection. Among available literature, there was no previous study that evaluated GzmB gene polymorphisms with the development of HCC. Alternatively, several studies have explored this association of these polymorphisms with other disorders and malignancies. In a recent Chinese study, Xu et al. [8] recruited 973 patients with vitiligo and 2,147 age- and gender-matched family-unrelated controls. It was discovered that having the rs8192917 C allele increased the likelihood of having vitiligo by around 40 %. The significant relationship of this SNP was identified in all three genetic models (co-dominance, dominance, and recessive).
In another study, Gaafar et al. investigated the role of GzmB gene polymorphism among 30 women with breast cancer and 38 other healthy Saudi Arabian women [9]. The study indicated that women carrying the C allele of rs8192917 polymorphism are more susceptible to the disease compared with those carrying the T allele. In a Dutch study, Hurkmans et al. investigated the role of rs8192917 in 347 patients with stage IV lung cancer under anti-programmed cell death (PD-1) therapy [10]. Patients with homozygous and heterozygous genotypes (TC and CC) had worse overall survival (OR: 1.60; 95%CI: 1.01 -2.52; p=0.044) and worse progression free survival (hazard ratio: 1.38; 95%CI:1.02 to 1.87; p=0.036) than those with wild types. In individuals infected with measles, rs8192917 was linked to subacute sclerosing panencephalitis, according to Yentur and his colleagues [11]. Corrales’s working team suggested that rs8192917 homozygous TT has linkage with improved kidney allograft outcomes [12]. In contrast, Espinoza et al. found that the CC genotype of rs8192917 in the patients who underwent bone marrow transplant was significantly associated with increased overall survival and reduced transplant-related mortality [13]. Several mechanisms have been postulated to explain this effect of rs8192917 as a risk for many diseases. This polymorphism is located in exon 2 of the gene. It involves a substitution of thymine with cytosine with a subsequent substitution of glutamine with arginine at amino acid number 55 of the polypeptide chain.
The most plausible explanation was proposed by McIlory et al. [14], who transfected primary glioblastoma cells with a mutant allele of this polymorphism and used x-ray crystallography to study the interaction between different amino acids in the GzmB structure. The study revealed that the presence of glutamine 55 in this peptide forms a hydrogen bond with the amino acid methionine 242. This interaction anchors the C-terminal α-helix to the surface of the protein, which seems to have a functional role in the catalytic properties of the enzyme. However, this interaction is disrupted when glutamine is substituted by arginine, and the protein loses some of its activity. In the same context, Oboshi et al. [15] have shown that rs8192917 can affect NK cell cytotoxicity. However, the authors did not explain the mechanism by which this polymorphism affects NK cells. These findings suggest that this polymorphism influences cytotoxic T cell function by altering the protein binding capacity with its substrate. In the present study, the haplotype CCG (C allele of rs7144366, C allele of rs8192917, and G allele of rs2236338) was significantly associated with an increased risk of HCC in patients with CHB. These alleles correspond to alanine, arginine, and histidine amino acid, respectively. This is in accordance with several previous reports that have investigated the three SNPs in different diseases. In a Korean study, Jeong et al. [16] investigated a total of 249 patients with non-segmented vitiligo (NSV) and 455 healthy controls to determine the impact of 5 SNPs (rs2236337, rs2236338, rs11539752, rs10909625, and rs8192917) in GzmB gene on patients’ susceptibility to this autoimmune disease. The haplotypes CGCCC consisting of rs2236337, rs2236338, rs11539752, rs10909625, and rs8192917 demonstrated a significant association with the disease. In another study, individuals presenting mutant haplotype of the three SNPs (rs7144366, rs8192917, and rs2236338) were 16 times more likely to have breast cancer as compared with those with wild type haplotype [9].
The mechanism by which the CCG haplotype increases the risk of HCC is not fully understood. Both mutant and wild type haplotypes of the alternative GzmB polypeptide isoforms are thought to have equal expression, stability, and proteolytic activity [14]. However, Gaafar et al. [9] found that CCG induced less apoptosis than TAA GzmB, while Sun et al. [17] found no difference. Other studies reported that the triple-mutated granzyme B variant encodes three amino acid changes with altered biological functions [18]. Alternatively, McIlory et al. [14] claimed that the three altered amino acids did not affect the active site or the substrate-binding cleft configuration. Furthermore, none of the three locations are involved in the active site or substrate-binding pocket. Additionally, the location of the triple mutant’s side chains revealed no steric incompatibility between the mutant side chains and the wild-type conformation. Apoptosis induction, in contrast, differed significantly between RAH and QPY GzmB. GzmB QPY (for glutamine, proline, and tyrosine, respectively) triggered apoptosis, as shown by the fact that caspase-3 is active in cell lysates and that dead cells let out LDH. But RAH GzmB (RAH stands for arginine, alanine, and histidine) did not cause caspase activity or cell death more than what was seen in control transfected cells. Three separate cell lines from different patients all showed the same results [14].
In the present study, The SNP rs8192917 is in tight link with rs2236338 (the measure D’ was 0.86). Furthermore, there was weak LD between rs2236338 and rs7144366 (the measure D’ was 0.57). Most previous studies indicated the presence of such a link, although with different degrees.
There is a significant degree of linkage disequilibrium between these mutations, as shown by many lines of evidence. Patients who are heterozygous for the Y245H variation are invariably heterozygous for the Q48R and P88A variants, while patients who were homozygous for the H245 variant were also homozygous for the R48 and A88 variants. Furthermore, cloning and sequencing of PCR products spanning exons 1–5 of the GzmB gene from a single heterozygous at all three sites confirmed that one chromosome coded for Q48 P88 Y245, whereas the other coded for R48 A88 H245 [14]. A study in Europe discovered a high level of LD among the three non-synonymous SNPs (rs8192917, rs11539752, and rs2236338) among European populations, resulting in different proteins [19,20]. Jin et al. [21] showed that rs8192917C was in a very strong LD with two other common non-synonymous SNPs, rs11539752 (r2 = 0.99) and rs2236338 (r2 = 0.93). In another study, the three SNPs had strong LD. In particular, rs8192917 had a stronger LD with rs11539752 (D′ = 0.951, r2 = 0.861) and rs2236338 (D′ = 0.942, r2 = 0.464,). A strong LD was also observed between rs8192917 and rs11539752 (D′ = 1.000, r2 = 0.549) [15].
CONCLUSIONS
Collectively, these data indicate that allele C of this SNP could be considered a risk factor for developing HCC. Furthermore, the haplotype block CCG (corresponding to the TC allele of rs7144366, the C allele of rs8192917, and the G allele of rs2236338) could also be considered a risk factor for the development of HCC in patients with CHB. In contrast, the haplotype block TAA of these SNPs has a protective role. There is a linkage disequilibrium between the three SNPs, especially between rs2236338 and rs8192917.
ACKNOWLEDGEMENT
The authors highly appreciate the great cooperation of included patients to complete the study. Also, we acknowledge the great help from all staff at Medical Research Unit/ College of Medicine/ Al-Nahrain University for their effort in molecular assays.
AUTHOR CONTRIBUTIONS
QSA conceptualized and supervised the experiment. QSA and HSA performed the experiment. HAS and FAA assisted in the sample collection and gross data recording and provided research assistance for data collection. QSA and HSA analyzed the data and interpreted the results. QSA, FAA and HAS drafted the manuscript and edited the manuscript. All authors agreed to be responsible for all elements of the work, including its accuracy and integrity.
CONFLICTS OF INTEREST
There is no conflict of interest among the authors.
References
- [1]Ferlay J, Shin HR, Bray F, Forman D, Mathers C, Parkin DM. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer. 2010 Dec 15;127(12):2893-917.
- [2]Bertoletti A, Tan AT. HBV as a target for CAR or TCR-T cell therapy. Curr Opin Immunol. 2020 Oct; 66:35-41.
- [3]Tan A, Koh S, Bertoletti A. Immune response in hepatitis B virus infection. Cold Spring Harb Perspect Med. 2015;5(8): a021428.
- [4]Lieberman J. Granzyme A activates another way to die. Immunol Rev. 2010;235(1):93-104.
- [5]Ben Safta T, Ziani L, Favre L, Lamendour L, Gros G, Mami-Chouaib F, et al. Granzyme B-activated p53 interacts with Bcl-2 to promote cytotoxic lymphocyte-mediated apoptosis. J Immunol. 2015 Jan 1;194(1):418-28.
- [6]Girnita DM, Webber SA, Brooks MM, Ferrell R, Girnita AL, Burckart GJ, et al. Genotypic variation and phenotypic characterization of granzyme B gene polymorphisms. Transplantation. 2009 Jun;87(12):1801-1806.
- [7]Mhaidat NM, Al-azzam SI, Alzoubi KH, Khabour OF, Gharaibeh BF. Granzyme B gene polymorphisms, colorectal cancer risk, and metastasis. J Cancer Res Ther. 2014 Jul-Sep;10(3):587-90.
- [8]Xu M, Liu Y, Liu Y. Li X, Chen G, Dong W, et al. Genetic polymorphisms of GZMB and vitiligo: A genetic association study based on Chinese Han population. Sci Rep. 2018;8, 13001
- [9]Gaafar A, Aljurf MD, Al Sulaiman A, Iqniebi A, Manogaran PS, Mohamed GE, et al. Defective gammadelta T cell function and granzyme B gene polymorphism in a cohort of newly diagnosed breast cancer patients. Exp Hematol. 2009; 37:838 48.
- [10]Hurkmans DP, Basak EA, Schepers N, Oomen-De Hoop E, Van der Leest CH, El Bouazzaoui S, et al. Granzyme B is correlated with clinical outcome after PD-1 blockade in patients with stage IV non-small-cell lung cancer. J Immunother Cancer. 2020 May;8(1): e000586.
- [11]Yentur SP, Aydin HN, Gurses C, Demirbilek V, Kuru U, Uysal S, et al. Granzyme B gene polymorphism associated with subacute sclerosing panencephalitis. Neuropediatrics. 2014;45, 309–313.
- [12]Corrales-Tellez E, Vu D, Shah T, Hutchinson I, Min DI. Association between granzyme B and perforin I polymorphisms and allograft outcomes in Hispanic kidney transplant recipients. Clin. Transpl. 2013;27: E308–E315.
- [13]Espinoza LJ, Takami A, Nakata K, Yamada K, Onizuka M, Kawase T, et al. Genetic variants of human granzyme B predict transplant outcomes after HLA matched unrelated bone marrow transplantation for myeloid malignancies. PLoS ONE. 2011:6: e23827
- [14]McIlroy D, Cartron PF, Tuffery P, Dudoit Y, Samri A, Autran B, et al. A triple mutated allele of granzyme B incapable of inducing apoptosis. Proc Natl Acad Sci USA. 2003; 100:2562 7.
- [15]Oboshi W, Watanabe T, Hayashi K, Nakamura T, Yukimasa N. QPY/RAH haplotypes of the GZMB gene are associated with natural killer cell cytotoxicity. Immunogenetics. 2018;70, 29–36.
- [16]Jeong KH, Kim SK, Seo JK, Shin MK, Lee MH. Association of GZMB polymorphisms and susceptibility to non-segmental vitiligo in a Korean population. Sci Rep. 2021;11(1):397.
- [17]Sun J, Bird CH, Thia KY, Matthews AY, Trapani JA, Bird PI. Granzyme B encoded by the commonly occurring human RAH allele retains pro-apoptotic activity. J Biol Chem. 2004; 279:16907-16911.
- [18]Zaitsu M, Yamamoto K, Ishii E, Teramura T, Nakadate N, Sako M, et al. High frequency of QPY allele and linkage disequilibrium of granzyme B in Epstein-Barr virus associated hemophagocytic lymphohistiocytosis. Tissue Antigens. 2004; 64: 611.
- [19]Guan, F, Niu Y, Zhang T, Liu S, Ma L, Qi T, et al. Two-stage association study to identify the genetic susceptibility of a novel common variant of rs2075290 in ZPR1 to type 2 diabetes. Sci Rep. 2016; 6,29586.
- [20]Guan F, Wei S, Zhang C, Zhang H, Zhang B, et al. A population-based association study of 2q32.3 and 8q21.3 loci with schizophrenia in Han Chinese. J Psychiatric Res. 2013; 47:712–717.
- [21]Jin Y, Birlea SA, Fain PR, Gowan K, Riccardi SL, Holland PJ, et al. Variant of TYR and autoimmunity susceptibility loci in generalized vitiligo. New Engl J Med. 2010; 362:1686–97.