Efficacy of soil-borne Burkholderia Cepacia-like MB-01 strain for shobicron and vertimec degradation
Abstract
Pesticides are necessary for agriculture, yet their highly toxic ingredients harm the ecosystem. Due to their toxicity, uncontrolled releases of large quantities of pesticides pollute the environment and provide a larger health risk to plants, animals, and humans. Bacteria are capable of degrading such pollutants and saving our ecosystem. In this study, a bacterial strain was isolated from Shobicron and Vertimec-treated lady’s finger soil using enrichment culture. The strain was identified as Burkholderia Cepacia-Like MB-01 based on morphological, physiological, and biochemical traits, as well as phylogenetic analysis of the 16S rRNA sequence. The bacterium grew best at 35 °C with a pH of 7. Furthermore, it was susceptible (S) to Cefepime and Penicillin but intermediate (I) resistant to Carbapenem and Tetracycline and resistant (R) to Ciprofloxacin, Kanamycin, and Gentamycin in an antibiotic sensitivity test. The rate of shobicron and vertimec degradation was measured over a five-day period using Mineral Salt (MS) medium. In its optimum growth condition, shobicron and vertimec degradation rates were around 76 % and 80 %, respectively. The isolated bacterial strain was capable of detoxifying shobicron and vertimec in the experiment. As a result, the bacterial strain could exploit as a possible shobicron and vertimec degrader for pesticide bioremediation.
INTRODUCTION
Bioremediation is a procedure that uses microorganisms or their enzymes to treat polluted places in order to restore them to their previous state [1]. Indeed, bioremediation is the act of breaking down harmful pollutants into less dangerous compounds using biological organisms. The bioremediation method involves a variety of live species. Majority of them are bacteria and fungus due to their quick growth rate and capacity to be genetically modified. To break down a pollutant, a widely used strategy for bioremediation involves activating naturally occurring microbial communities and providing them with nutrients and other needs. We should utilize bioremediation to clean soil, water, and other materials since it is less damaging and poses fewer risks to the environment [2].
Pesticides are a group of poisonous compounds used mostly in agriculture. A pesticide is a mixture of substances used to eliminate or control a pest, which can include vectors of human or animal disease, undesirable plant species, or animals that cause damage to agricultural food products, timber and timber products, or animal feed during production, processing, storage, or marketing. Thousands of pesticide formulations exist around the world, containing almost 1500 registered ingredients [3]. More than 55 % of the area used for agricultural production in developing countries uses about 26 % of pesticides produced worldwide [4]. Farmers in Bangladesh frequently use various pesticide models on their agricultural areas, typically on the advice of unskilled vendors [5]. Pesticides are applied in granules, liquids, and powders. Depending on the attacking pests in Bangladesh, farmers have utilized pesticides such as cypermthrin, carbofuran, amcozinon, profenofos, abamectin, and malathion.
Hazardous pesticides can have acute and/or chronic poisoning effects, putting children at danger, and are recognized as a worldwide concern. In many places of the world, their widespread use has resulted in health concerns and deaths, typically as a result of occupational exposure and unintentional poisonings. Agricultural and public health professionals are exposed to the most dangerous pesticides due to their handling, mixing, and application. Pesticide residues in food and, possibly, drinking water can be a concern for the general public [6].
Agricultural, residential, and industrial wastes are the main contributors of water contamination. When sewage is dumped into rivers, it is the most contaminant of fresh water. The amount of dissolved oxygen in the water decreases, and the Biological Oxygen Demand (BOD) decreases. When large amounts of fertilizer, herbicides, and pesticides are washed into rivers by rain, they pose a major threat to human life [7]. Shobicron is a brand name for two ingredients: cypermethrin and profenofos. It’s used to fight pests like fruit flies, white flies, and dragonflies in a variety of agricultural crops like brinjal, lady’s finger, cotton, guava, mango, and banana. Cypermethrin is a synthetic pyrethroid that is widely used as a pesticide in agriculture, household, and animal husbandry, primarily to treat cracks, crevices, and spots for insect control [8]. Cypermethrin has a very poor water solubility [9]. It moves from a liquid solution to an adjourned molecule because it is hydrophobic [10]. The urine excretion of cypermethrin metabolites in humans was completed 48 hours after the last of five daily 1.5 mg doses [11]. Profenofos is an insecticide containing organophosphates. It’s a pale yellow to amber-colored liquid with a garlic-like odor [12]. The resistance of the acetylcholinesterase enzyme is the mechanism of action of profenofos. Although it is utilized as a racemate, the S (-) isomer is a far more powerful inhibitor. Profenofos can prevent the cholinesterase enzyme from working properly. This can lead the neurological system to become over stimulated, resulting in nausea, dizziness, and confusion. Higher doses of profenofos can cause even more severe side effects, including respiratory paralysis and death. Profenofos has also been shown to affect bees and fish. Vertimec, as a brand name, is a popular insecticide in Bangladesh, where it is used on a wide range of crops. Abamectin is the active component. It is employed in a variety of agricultural crops, including brinjal, lady’s finger, tea, jujube, litchi, and others, to combat pests such as red spinder mite and mite. Abamectin is widely used as an insecticide in agriculture and as an anti-parasitic medication in veterinary medicine, and it has resulted in significant environmental contamination by posing a risk to non-target soil invertebrates and aquatic systems. Avermectin is made up of two homologues: avermectin B1a and avermectin B1b [13]. Abamectin is extremely poisonous to aquatic invertebrates and highly toxic to fish. As a result, abamectin should not be disposed of in water. Abamectin contaminates groundwater and harms the environment.
Thus, the present study was carried out to isolate and characterize shobicron and vertimec degrading bacteria from lady’s finger field soil, as well as determine their degradation efficiency for further exploitation of the strain for potential bioaugmentation.
MATERIALS AND METHODS
Sample collection
One gram of shobicron and vertimec-treated soil sample was collected from agricultural fields in Budpara and Rajshahi, Bangladesh that had a 5-7-years history of shobicron and vertimec use. Soil sample was collected and employed as a source of inoculums for enrichment culture. This research work was carried out over the course of seven months, from May to December 2019. The entire project was carried out in the microbiology laboratory, Department of Genetic Engineering and Biotechnology of the University of Rajshahi, Bangladesh.
Cytotoxicity analysis of shobicron and vertimec using Artemia salina through LC50
The brine shrimp lethality bioassay is a useful technique for toxicity screening, heavy metal detection, pesticide detection, fungal toxins detection, and dental cytotoxicity testing [14]. Brine shrimp eggs were cultured in artificial seawater to obtain nauplii. The nauplii were counted and placed in test tubes visually. Then, in three replications, different amounts of shobicron and vertimec were delivered to tubes, with control doses kept at the same level as before. To assess the percentage of survived nauplii after 12, 24, 36, and 48 hours, data was collected and statistically analyzed using probit mortality software.
Enrichment of shobicron and vertimec degrading bacteria
One gram of shobicron and vertimec-treated soil was suspended in 100 mL of mineral salts (MS) medium (Himedia, India) supplemented with shobicron and vertimec in separate 250 mL Erlenmayer flasks (Eisco lab, India). To account for potential abiotic disappearance of shobicron and vertimec, control flasks without an inoculum were also prepared. Shaking at 120rpm for 3 days on an orbital shaker was the principal enrichment method. The cultures were employed as a source of inoculum in the following tests once they acquired sufficient turbidity.
By plating out on mineral salts agar medium containing shobicron and vertimec, microorganisms capable of decomposing shobicron and vertimec were identified from enrichment cultures. The plates were incubated at 37 °C for 2 to 3 days, and one strain that grew on the medium and had a greater degrading ability was named MB-01 and stored for future use.
Morphological, and biochemical characterization of the isolate
After 24 hours of growth on MS agar at 37 °C, the colony features of the isolated bacteria (colony form, color, margin, opacity, consistency, elevation) were studied under an inverted microscope (LABOMED CxL). The morphological (Gram staining and motility tests) and biochemical (catalase, oxidase, TSI, methyl red, lactose fermentation tests, carbohydrates such as glucose, mannose, arabinose, sucrose, lactose, and cellulose intake) characterizations were carried out according to the protocol [15].
Antibiotic sensitivity test
The antibiotic sensitivity and resistance pattern of the isolated bacteria were determined using the Kirby- Bauer disc diffusion method [16]. For the antibiotic sensitivity test, the isolated bacterial strain was cultured overnight in nutrient broths using a shaker at 37 °C and 160 rpm. The LB (Luria Bertani) agar medium (Himedia, India) was prepared first, and then the sterile liquid medium was dispersed in sterile petridishes for preparing culture plates. Each petridish received approximately (15-20) ml of the media and was placed in the laminar airflow cabinet to solidify. Then, for the isolate, overnight grown LB culture (OD = 0.5) was poured into the nutrient agar plate and dried. Antibiotic disks (Table 1) were prepared using commercially available and regularly prescribed antibiotics, which were inserted in the center of the plates and incubated overnight at 37 °C. After an overnight incubation period, the zones on the plate were observed and quantified using a millimeter scale.
Effect of pH and temperatures on bacterial growth
The media were prepared, and pH adjusted to a range of 5 to 8 on a scale of 1 to 8. The bacteria were then inoculated in a medium that had already been prepared in front of the laminar airflow cabinet. After the culture was inoculated, it was placed in an incubator to maintain the various temperature ranging from 25 °C to 45 °C with 5 °C interval to test their growth efficiency. By using a spectrophotometer (Wincom, China) to measure optical density at 600nm, the growth rates were recorded over time.
PCR amplification and sequencing of 16S rRNA gene and phylogenetic analysis
The Tiangen DNA extraction kit was used to extract the genomic DNA of the isolated shobicron and vertimec degrading bacterium. The 16S rRNA genes were amplified using forward primer in a polymerase chain reaction (PCR) (27 F – AGA GTT TGA TCM TGG CTC AG and 1492 R – GGT TAC CTT GTT ACG ACT T). Invitrogen (Thermo fisher scientific, Bangladesh) provided all the PCR reagents. The PCR reactions were carried out in a thermal cycler (Applied Biosystem 9700) with the following amplification conditions: initial denaturation at 95 °C for 5 minutes, followed by 30 cycles of denaturation at 94 °C for 1 minute, annealing at 55 °C for 1 minute, extension at 72 °C for 1 minute, and final extension at 72 °C for 10 minutes. The Freeze ‘N Squeeze DNA Gel Extraction Spin Columns (Bio-Rad, India) were used to purify the PCR results, which were then sequenced on both strands on the genetic analyzer (Prism 310). Chromas, a bioinformatics program, were used to alter the sequences. The BLASTN (http://www.ncbi.nih.gov/BLAST) algorithm was used to compare the 16S rRNA gene sequences to the 16S rRNA gene sequences of other organisms. NCBI (National center for biotechnology information) BLAST (www.ncbi.nlm.nih.gov/Blast) was used to create the phylogenetic tree.
Determination of shobicorn and vertimec degradation rate
The isolate was grown in a 150 mL conical flask with 100 mL Mineral salts medium (MSM) supplemented with shobicron and vertimec to test its growth. A spectrophotometer was used to calculate the ability to the isolate to degrade pesticides by measuring turbidity at 600 nm. To compare the deterioration of the isolate, a control flask was utilized without pesticides. Control and isolates had their ideal density values evaluated and recorded.
RESULTS
Cytotoxicity test of the 99% pure shobicron and vertimec
The toxic effects of pure shobicron and vertimec on A. salina were investigated using a brine shrimp lethality assay. After only 12 hours of exposure, the pure shobicron and vertimec LC50 were 10.102 g/ml and 9.871 g/ml, respectively, according to probit analysis, and no live nauplii remained after 48 hours (Table 1). Shobicron and vertimec exhibited a significant mortality effect on nauplii, as evidenced by visual detection.
Table 1. Determination of LC50 value of the shobicron and vertimec against Artemia salina after 12, 24, 36, and 48 h of exposure.
Isolation and screening of shobicron and vertimec degrading bacteria
The bacteria that degrade shobicron and vertimec were identified using 99% pure shobicron and vertimec at 2 µg/ml and 4 µg/ml, respectively. The shobicron and vertimec-enriched MS plates were incubated at 37 °C for 24 hours, and bacterial colonies were observed to develop on the medium containing the aforesaid concentrations of shobicron and vertimec (Figure 1a), with no growth at concentrations higher than that.
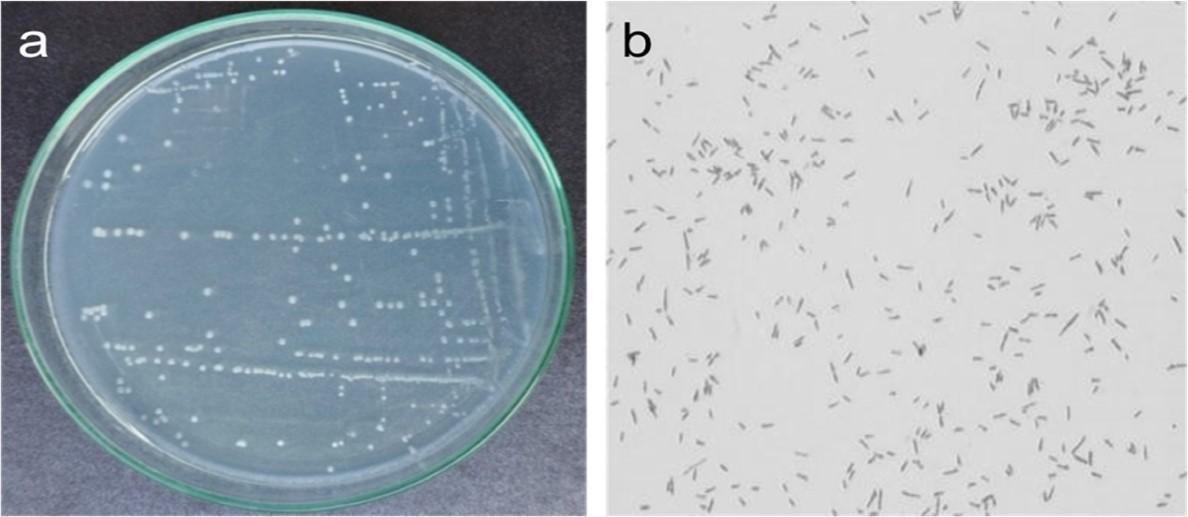
Morphological and biochemical characterization
The strain formed a smooth colony with an average diameter of 0.6–1 mm in biochemical tests. The cells were straight rods with rounded ends, measuring 0.4–0.7 lm in width and 1.2–1.7 lm in length. It was a gram-negative bacterium (Figure 1b). The strain was positive for oxidase, nitrate reduction, and beta-glactocidase, but not for arginine dihydrolase, beta-glucosidase (aesculin hydrolysis), protease (gelatin liquefaction), tryptophanase, or urease activity. L-arabinose, D-glucose, D-mannose, and D-mannitol were among the carbohydrates that the bacterium could use.
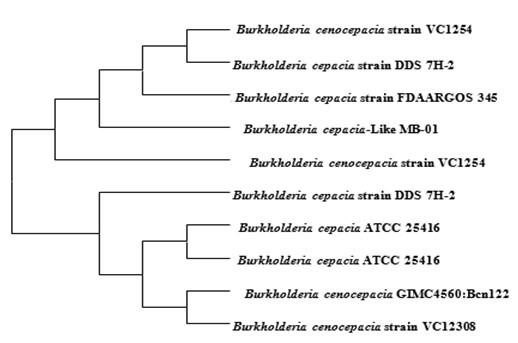
To identify the isolated bacterium, the 16S rRNA gene was amplified by PCR and sequenced from Invent technologies, Bangladesh. The strain MB-01 demonstrated very high similarity with the species of genus Burkholderia, and more than 99% gene homology was observed with Burkholderia cepecia complex (Bcc) species. Phylogenetic analysis of 16S rRNA gene also clustered the strain GB-01 within the clade of Burkholderia cepecia complex (Bcc) species in phylogenetic tree (Figure 2). On the basis of morphological, physiological and biochemical properties, combined with 16S rRNA gene sequence analysis, the strain MB-01 was identified as Burkholderia cepacia-like species.
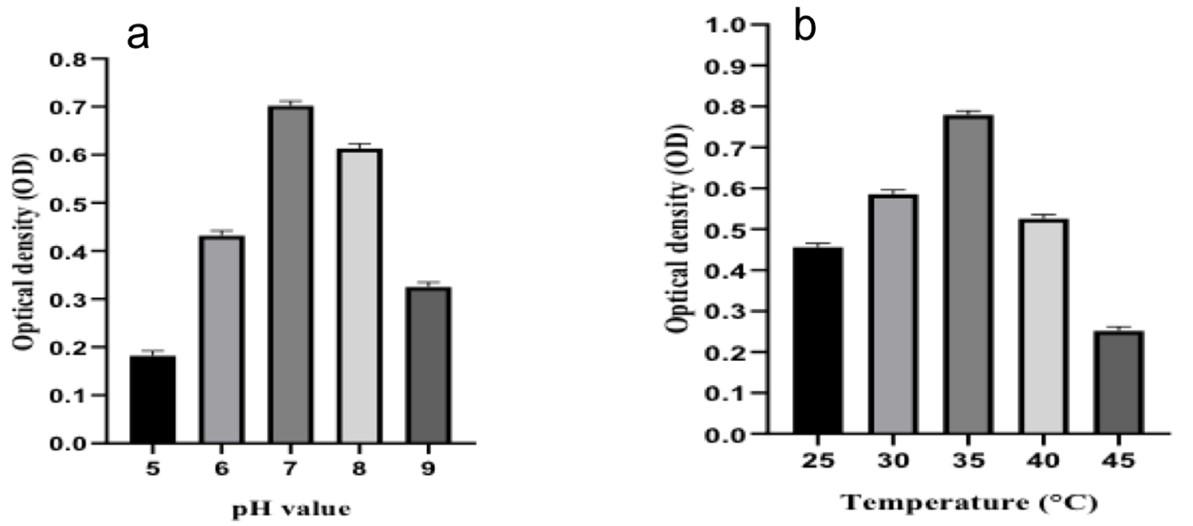
Physiological characterization
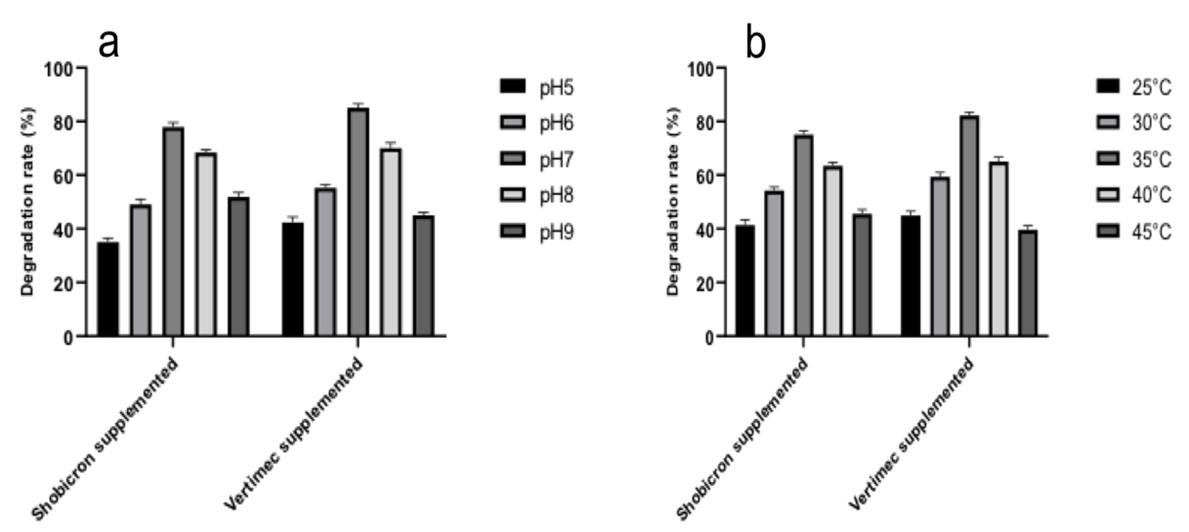
This test was carried out to assess the optimum pH and temperature for bacterial growth. The strain was grown on MS media at temperatures of 25 °C, 30 °C, 35 °C, 40 °C, and 45 °C in a pH range of 5 to 9. After measuring OD at 600nm in a Spectrophotometer, it was observed that the strain’s optimum growth was best at pH 7 (Figure 3a) and 35 °C (Figure 3b).
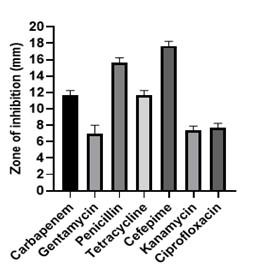
Antibiotic sensitivity test
The result showed that Burkholderia Cepacia-Like MB-01 was susceptible (S) to Cefepime and Penicillin but intermediate (I) resistant to Carbapenem, and Tetracycline and resistant (R) to Ciprofloxacin, Kanamycin, and Gentamycin (Figure 4).
Shobicron and vertimec degrading efficiency of Burkholderia Cepacia-like MB-01
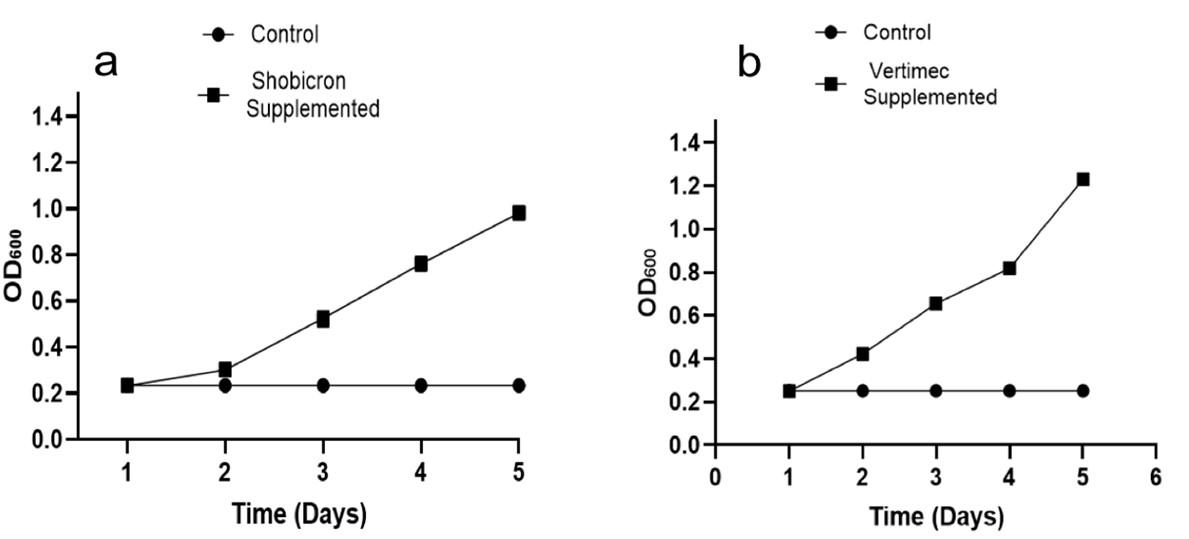
The strain MB-01 was cultivated on shobicron and vertimec supplemented liquid Mineral salts medium (MSM) at 35 °C for 5 days in order to gain relative degradation. The strain decomposed shobicron and vertimec, as evidenced by a steady increase in turbidity (OD600) over a 5-day period (Figure 5). When shobicron and vertimec were employed as the only source of carbon, the strain’s growth was dramatically boosted when compared to the control. Over 5 days of incubation at 35 °C, strain MB-01 showed a clear increase in growth using shobicron and vertimec, with a 76 %, and a 80 % rise in OD600, respectively.
To be more specific, we ran this experiment at different pH and temperature levels to see how well it degraded, and we obtained comparable results (Figure 6) as shown in Figure 5.
DISCUSSION
Biological processes like microbial decomposition are considered a safe and effective way to remove contaminants from the environment [17, 18]. Bioremediation is closely linked to the decomposition, or detoxification, of numerous pollutants and harmful compounds in the environment [19, 20]. Some pesticides are quickly degraded by microorganisms, while others are harmful. Pesticides are metabolized by a variety of bacteria, including members of the genera Alcaligenes, Flavobacterium, Pseudomonas, and Rhodococcus. Microbial degradation is influenced by a wide range of environmental conditions in addition to the presence of bacteria. Pesticide degradation is influenced by a variety of environmental parameters such as pH, temperature, and nutrients [21, 22]. Pesticides can be broken down by microorganisms that use them as carbon and energy sources in specific ways [23].
Bioremediation is a critical method for reducing pollution and restoring contaminated soil and water. We should utilize bioremediation to clean soil, water, and other materials since it is less damaging and poses fewer risks to the environment [24]. Because of its environmental stability, toxicity, and biological reservation in organisms, pesticides cause serious health problems. Due to a lack of sufficient understanding or a desire to maximize their profit, Bangladeshi farmers are currently using pesticides in quantities that are 10-15 times higher than those recommended by the government. In general, Bangladeshi farmers do not follow the recommended dosages and apply pesticides in excessive amounts [25]. Pesticide residues accumulate in the food chain, drinking water, and the environment as a result of this excessive application [26]. It poses a significant health risk to present and future generations [27]. In this study, a soil sample with a history (5-7 years) of shobicron and vertimec use was used to isolate a bacterial strain. We hypothesized that shobicron and vertimec would be digested and broken down by the bacteria.
The toxicity of a pesticide is assessed by administering different doses of the active ingredient (a.i.) and each of its manufactured products to test animals. The active ingredient is the pesticide’s chemical component that controls the pest. The toxicities of different active components and different formulations containing the same active ingredient can be compared using LD50 and LC50 values. The lower the LD50 or LC50 of a pesticide, the more hazardous it is to people and animals. As a result, we first looked at the toxicity profiles of the pesticides on Artemia salina. After 36 hours of exopuse, the LC50 values for Shobicron and vertimec were calculated to be 2.454323 g/ml and 2.210981 g/ml, respectively, and are considered extremely toxic to humans and animals.
We used toxic shobicron and vertimec as a sole carbon source for the isolated bacteria to test detoxification efficiency. Research indicated that certain type of bacterial species have the capability to exploit sulfadoxine [28], organophophorus [29], tributyl phosphate [30], chlorpyrifos [31], and organophosphorous [32] pesticides as carbon and energy source. In this study, morphological and biochemical tests were also carried out, and the isolated strain was gram negative, rod shaped, as previously identifying characteristics of burkholderia genus [33]. Burkholderia is a highly versatile bacterium, which is capable of degradation of many hazardous pollutants [34, 35], and it has a role in protection and growth promotion of plants [36, 37].
In this research, penicillin, kanamycin, gentamycin, tetracycline, ciprofloxacin, cefepime, carbapenem were used to test antibiotic sensitivity of the isolated bacteria. The result showed that strain was susceptible (S) to Cefepime and Penicillin but intermediate (I) resistant to Carbapenem, & Tetracycline and resistant (R) to Ciprofloxacin, Kanamycin, and Gentamycin. Previous research suggested that the strain resistant to gentamycin and ciprofloxacin antibiotics; and were sensitive to penicillin, cefepime [38, 39].
Bacteria that degrade pesticides have been found so far, and they comprise both Gram positive and Gram negative bacteria [40]. Only a gram-negative bacterium was identified in this study. Atikur et al. found that two bacterial strains, Achromobacter and Diaphorobacter, can biodegrade pesticides [41]. This was backed up by other scientists [42, 43]. The majority of previously detected bacteria could only digest one pesticide. However, in this case, we identified one strain that could breakdown two pesticides. In its ideal state, this strain could breakdown both pesticides with an over 75% degradation rate. Similar results (Carbofuran degradation rate of about 80%) were obtained by Ekram et al. [44]. As a result, this research has a lot of promise in terms of pesticide bioremediation. The current study was carried out in a laboratory setting. As a result, Burkholderia Cepacia-Like MB-01 should be evaluated with industrial waste and in natural habitats in future investigations. Furthermore, before being used to detoxify pesticides in the environment, Burkholderia Cepacia-Like MB-01 should be examined for pathogenicity to aquatic organisms like as fish, prawns, and crab [45, 46].
CONCLUSION
Environmental pollution is being brought on by Bangladesh’s massive rise in pesticide use. Reducing the amount of pesticide in the environment may be accomplished by bioremediation. The effectiveness of Burkholderia Cepacia-Like MB-01 in biodegrading shobicron and vertimec is revealed by this study, and it may be helpful for bioremediation programs aimed at enhancing soil quality. This study will enable many researchers to explore the crucial aspects of employing the natural isolate Burkholderia Cepacia-Like MB-01 for shobicron and vertimec biodegradation. As a result, a novel theory about the use of Burkholderia Cepacia-Like MB-01 to clean up pesticide-contaminated soil can develop.
ACKNOWLEDGMENT
None
AUTHOR CONTRIBUTIONS
MMZ, MMJ, MMM, SI, MJP, JB, AI: Methodology, Validation, Formal analysis, Investigation, Data curation, Writing–original draft. MSU, MAS: Conceptualization, Validation, Resources, Data curation, Project administration, Editing. SZ: Supervision, Funding acquisition.
CONFLICTS OF INTEREST
There is no conflict of interest among the authors.
References
- [1]Glazer AN, Nikaido H. Microbial biotechnology: fundamentals of applied microbiology. Cambridge University Press; 2007.
- [2]Rani K, Dhania G. Bioremediation and biodegradation of pesticide from contaminated soil and water—a noval approach. Int J Curr Microbiol App Sci. 2014; 3(10): 23-33.
- [3]Zaller JG. Daily Poison: Pesticides-an Underestimated Danger. Springer Nature; 2020.
- [4]Singh B. Role of pesticides in management of crop pests. Jodhpur: Scientific Publishers (India); 2012.
- [5]Alam MK. Determination of cypermethrin, chlorpyrifos and diazinon residues in tomato and reduction of cypermethrin residues in tomato using rice bran. World. 2013; 1(2): 30-5.
- [6]World Health Organization. Preventing disease through healthy environments: exposure to lead: a major public health concern. World Health Organization. License: CC BY-NC-SA 3.0 IGO. 2019
- [7]Khan WA, Ali S, Shah SA. Water Pollution: Sources and Its Impact on Human Health, Control and Managing. Journal of International Cooperation and Development. 2022;5(1): 69-69.
- [8]Patel P, Prajapati R. Possible Toxicological Effect of Pesticides, Cypermethrin on Earthworm (Esiania fetida). International Journal of Research in Engineering, Science and Management. 2020; 3(12): 138-43.
- [9]Korkmaz V, Güngördü A, Ozmen M. Comparative evaluation of toxicological effects and recovery patterns in zebrafish (Danio rerio) after exposure to phosalone-based and cypermethrin-based pesticides. Ecotoxicology and Environmental Safety. 2018; 160: 265-72.
- [10]Singh RP, Singh R. Adsorption and movement of cypermethrin on Indian soils amended with cationic, non-ionic and anionic surfactants. Adsorption Science & Technology. 2004; 22(7): 553-64.
- [11]Al-Autaish HHN. Clinical study on toxicity of cypermethrin in Arrabi sheep. Doctoral dissertation, M. Sc. Thesis, Coll. of Veterinary Medicine, Univ. of Basrah. 2010; pp: 76.
- [12]Guo J, Tong M, Tang J, Bian H, Wan X, He L, et al. Analysis of multiple pesticide residues in polyphenol-rich agricultural products by UPLC-MS/MS using a modified QuEChERS extraction and dilution method. Food chemistry. 2019; 274: 452-9.
- [13]Ali SW, Li R, Zhou WY, Sun JQ, Guo P, Ma JP, et al. Isolation and characterization of an abamectin-degrading Burkholderia cepacia-like GB-01 strain. Biodegradation. 2010; 21(3): 441-52.
- [14]Suryawanshi V, Yadav A, Mohite S, Magdum CS. Toxicological Assessment using Brine Shrimp Lethality Assay and Antimicrobial activity of Capparis Grandis. Journal of University of Shanghai for Science and Technology. 2020; 22(11): 746-59.
- [15]Lyne PM, Grange JM. Collins and Lyne’s microbiological methods. 8th Edition, Oxford University Press, New York. 2004.
- [16]Bauer AW. Antibiotic susceptibility testing by a standardized single disc method. Am J clin pathol. 1966; 45: 149-58.
- [17]Bwapwa JK. Factors affecting the bioremediation of industrial and domestic wastewaters. In Microbial Biodegradation and Bioremediation, Elsevier. 2022; (pp. 461-472).
- [18]Singh R. Microorganism as a tool of bioremediation technology for cleaning environment: a review. Proceedings of the International Academy of Ecology and Environmental Sciences. 2014; 4(1): 1.
- [19]Harms H, Schlosser D, Wick LY. Untapped potential: exploiting fungi in bioremediation of hazardous chemicals. Nature Reviews Microbiology. 2011; 9(3): 177-92.
- [20]Hossain K, Ismail N. Bioremediation and detoxification of pulp and paper mill effluent: A review. Research Journal of Environmental Toxicology. 2015; 9(3): 113.
- [21]Singh BK, Walker A, Wright DJ. Bioremedial potential of fenamiphos and chlorpyrifos degrading isolates: influence of different environmental conditions. Soil Biology and Biochemistry. 2006; 38(9): 2682-93.
- [22]Md Z, Mohammed A, Badrul I, Azza SA, Mohamed AE, Arif A. Bioremediation: A tool for environmental cleaning. African Journal of Microbiology Research. 2009; 3(6): 310-4.
- [23]Huang Y, Xiao L, Li F, Xiao M, Lin D, Long X, et al. Microbial degradation of pesticide residues and an emphasis on the degradation of cypermethrin and 3-phenoxy benzoic acid: a review. Molecules. 2018; 23(9): 2313.
- [24]Hussain A, Rehman F, Rafeeq H, Waqas M, Asghar A, Afsheen N, et al. In-situ, Ex-situ, and nano-remediation strategies to treat polluted soil, water, and air–A review. Chemosphere. 2022; 289: 133252.
- [25]Chowdhury MG, Rahman MA, Miaruddin M, Khan MH, Rahman MM. Assessment of pesticides and ripening chemicals used in selected vegetables at different locations of Bangladesh. Bangladesh Journal of Agricultural Research. 2019; 44(2): 261-79.
- [26]Liu L, Zheng X, Wei X, Kai Z, Xu Y. Excessive application of chemical fertilizer and organophosphorus pesticides induced total phosphorus loss from planting causing surface water eutrophication. Scientific Reports. 2021; 11(1): 1-8.
- [27]Rahman S. Farm-level pesticide use in Bangladesh: determinants and awareness. Agriculture, ecosystems & environment. 2003; 95(1): 241-52.
- [28]Zhang W, Xu D, Niu Z, Yin K, Liu P, Chen L. Isolation and characterization of Pseudomonas sp. DX7 capable of degrading sulfadoxine. Biodegradation. 2012; 23(3): 431-9.
- [29]Chino-Flores C, Dantán-González E, Vázquez-Ramos A, Tinoco-Valencia R, Díaz-Méndez R, Sánchez-Salinas E, et al. Isolation of the opdE gene that encodes for a new hydrolase of Enterobacter sp. capable of degrading organophosphorus pesticides. Biodegradation. 2012; 23(3): 387-97.
- [30]Ahire KC, Kapadnis BP, Kulkarni GJ, Shouche YS, Deopurkar RL. Biodegradation of tributyl phosphate by novel bacteria isolated from enrichment cultures. Biodegradation. 2012; 23(1): 165-76.
- [31]Harishankar MK, Sasikala C, Ramya M. Efficiency of the intestinal bacteria in the degradation of the toxic pesticide, chlorpyrifos. 3. Biotech. 2013; 3(2): 137-42.
- [32]Kanekar PP, Bhadbhade BJ, Deshpande NM, Sarnaik SS. Biodegradation of organophosphorus pesticides. Proceedings-Indian National Science Academy Part B. 2004; 70(1): 57-70.
- [33]AuCoin DP, Crump RB, Thorkildson P, Nuti DE, LiPuma JJ, Kozel TR. Identification of Burkholderia cepacia complex bacteria with a lipopolysaccharide-specific monoclonal antibody. Journal of medical microbiology. 2010; 59(Pt 1): 41.
- [34]Morya R, Salvachúa D, Thakur IS. Burkholderia: an untapped but promising bacterial genus for the conversion of aromatic compounds. Trends in Biotechnology. 2020; 38(9): 963-75.
- [35]Seo JS, Keum YS, Li QX. Bacterial degradation of aromatic compounds. International journal of environmental research and public health. 2009; 6(1): 278-309.
- [36]Zaman NR, Chowdhury UF, Reza RN, Chowdhury FT, Sarker M, Hossain MM, et al. Plant growth promoting endophyte Burkholderia contaminans NZ antagonizes phytopathogen Macrophomina phaseolina through melanin synthesis and pyrrolnitrin inhibition. PloS one. 2021; 16(9): e0257863.
- [37]Kong P, Hong C. Endophytic Burkholderia sp. SSG as a potential biofertilizer promoting boxwood growth. PeerJ. 2020; 8: e9547.
- [38]Roy P, Ahmed N, Biswal I, Grover RK. Antimicrobial susceptibility pattern of Burkholderia cepacia isolates from patients with malignancy. Journal of Global Infectious Diseases. 2014; 6(2): 90.
- [39]Moore JE, Crowe M, Shaw A, McCaughan J, Redmond AO, Elborn JS. Antibiotic resistance in Burkholderia cepacia at two regional cystic fibrosis centres in Northern Ireland: is there a need for synergy testing?. Journal of Antimicrobial Chemotherapy. 2001; 48(2): 319-21.
- [40]DeLorenzo ME, Scott GI, Ross PE. Toxicity of pesticides to aquatic microorganisms: a review. Environmental Toxicology and Chemistry: An International Journal. 2001; 20(1): 84-98.
- [41]Rahman MA, Arefin AS, Saha O, Rahaman MM. Isolation and identification of pesticides degrading bacteria from farmland soil. Bangladesh Journal of Microbiology. 2018; 35(2): 90-4.
- [42]Ren HF, Li SQ, Liu SJ, Liu ZP. Isolation and characterization of a p-chloroaniline-degrading bacterial strain. Huan Jing ke Xue= Huanjing Kexue. 2005; 26(1): 154-8.
- [43]Xia ZY, Zhang L, Zhao Y, Yan X, Li SP, Gu T, et al. Biodegradation of the herbicide 2, 4-dichlorophenoxyacetic acid by a new isolated strain of Achromobacter sp. LZ35. Current microbiology. 2017; 74(2): 193-202.
- [44]Ekram MA, Sarker I, Rahi MS, Rahman MA, Saha AK, Reza MA. Efficacy of soil‐borne Enterobacter sp. for carbofuran degradation: HPLC quantitation of degradation rate. Journal of basic microbiology. 2020; 60(5): 390-9.
- [45]Ortiz-Hernández ML, Sánchez-Salinas E, Dantán-González E, Castrejón-Godínez ML. Pesticide biodegradation: mechanisms, genetics and strategies to enhance the process. Biodegradation-life of Science. 2013; 10: 251-87.
- [46]Ravi RK, Bhawana P, Fulekar MH. Bioremediation of persistent pesticides in rice field soil environment using surface soil treatment reactor. International Journal of Current Microbiology and Applied Sciences. 2015; 4(2): 359-69.