Detection of multidrug resistant Salmonella spp. from healthy and diseased broilers having potential public health significance
Abstract
Multidrug resistant (MDR) Salmonella spp. poses significant global public health concern by causing food-borne infections. This study aimed to detect MDR Salmonella spp. from healthy and diseased broiler chickens in the Mymensingh and Jamalpur districts of Bangladesh. Total 70 samples comprising feces (n=20), chicken meat (n=30), and visceral organs i.e. liver, lung, and kidney (n=20) were collected. Salmonella were isolated and identified by culture, biochemical tests and PCR. The antibiogram study was performed by the disk diffusion method. By PCR, 30% (21/70; 95% CI: 19.32-40.05%) samples were positive for Salmonella spp., of which significantly (p=0.005) higher occurrence were detected in feces (50%; 95% CI: 29.93-70.07%) compared to chicken meat (10%; 95% CI: 3.46-25.62%) and visceral organs (40%; 95% CI: 21.88-61.34%). By antibiogram, all the Salmonella isolates were resistant to amoxicillin, and frequently (90.48-19.05%) resistant to tetracycline, ceftazidime, chloramphenicol, colistin, and ciprofloxacin. The significantly higher resistance of chloramphenicol, tetracycline, and ceftazidime were observed in the internal organs of broilers. Interestingly, 80.95% (17/21; 95% CI: 59.99-92.33%) Salmonella isolates were MDR in nature. The range of multiple antibiotic resistance (MAR) index of Salmonella isolates varied from 0.29 to 0.86. The high occurrence of MDR and MAR Salmonella in broilers detected in our present study could reveal a high risk to public health and these organisms could be transmitted to humans through the food supply. We suggest that effective prevention and control measures should be implemented to reduce their potential contamination and to minimize the emergence of antibiotic resistance.
INTRODUCTION
Poultry farming has become a profitable and dependable agricultural business in Bangladesh. In addition, it plays a momentous role in the employment generation and economic growth of Bangladesh [1]. Poultry provides additional income to rural people [2]. Furthermore, poultry delivers about 37% of the total meat supply to the people of Bangladesh and covers more than 12.5% of total daily proteins per capita [3]. But the entry of different infectious diseases e.g. salmonellosis, avian colibacillosis, mycoplasmosis, fowl cholera, avian influenza, Newcastle disease, infectious bronchitis, aspergillosis, and others hinder the further advancement of poultry production [4]. Among them, multidrug resistant (MDR) Salmonella spp. are deemed as major botherations in the uplifting of Bangladesh’s poultry sector by causing drastic poultry illness and deaths annually [5].
Salmonella spp. is one of the most frequently isolated foodborne pathogens that develops approximately 153 million enteric diseases and 155,000 deaths per year globally [6-8]. In poultry, Salmonella spp. is devastating for developing avian salmonellosis, increasing mortality rates, and reducing hatchability and fertility rates [3]. Poultry products especially meat and eggs play a pivotal role in Salmonella contamination. Incidences of food-poisoning diseases triggered by these pathogens have been increasing remarkably in the last several years. In human, Salmonella spp. cause human salmonellosis. Poultry-originated foods are thought to be the main reasons for human salmonellosis, as poultry especially broilers are important reservoirs of Salmonella spp. [9]. In addition, poultry and poultry-originated foods generally act as crucial sources for the sporadic outbreaks of human salmonellosis globally. As Salmonella spp. are naturally gut-originated pathogens in poultry, the food supply chain makes an important scope for the transmission of Salmonella infections to humans [10].
Antimicrobial resistance (AMR) is considered a worldwide health problem jeopardizing all one-health components [11]. The indiscriminate use of antibiotics triggers selection pressure and develops antibiotic resistance in bacteria [12]. In addition, antibiotics are being used as growth promoters in modern poultry, especially broiler production that also triggers the development of AMR in poultry. Globally, the speculation deaths due to AMR consequences will be more than 300 million per year, if significant steps won’t be taken by 2050 [13]. The world critics have warned that the low- and middle-income countries will face the worst impacts of AMR. According to the world health organization, Bangladesh is at high risk of AMR consequences [14].
The detection of MDR Salmonella spp. from broilers was previously recorded in Bangladesh [5, 15, 16]. However, it needs regular surveillance to determine the actual prevalence of Salmonella in broilers in Bangladesh. Therefore, the present study was carried out to detect MDR Salmonella in both healthy (feces, and meat) and diseased (visceral organs) broiler samples.
MATERIALS AND METHODS
Sample size calculation
The sample size of our present study was calculated following by the prevalence of Salmonella spp. (23.53%) isolated from broilers in Bangladesh [16]. The formula we followed for the sample size calculation was described previously [17]: n = Z2pq/d2, where, n = desired sample size, Z = the standard normal deviation (1.96 at 95% confidence level), p = prevalence (23.53% or 0.2353), q = 1-p = 1-0.2353 = 0.7647, d = precision at 10% (d = 0.1). So, n= (1.96)2×0.2353×0.7647/ (0.1)2 = 69.123. Therefore, we collected 70 samples from broiler chickens.
Sampling site and sampling
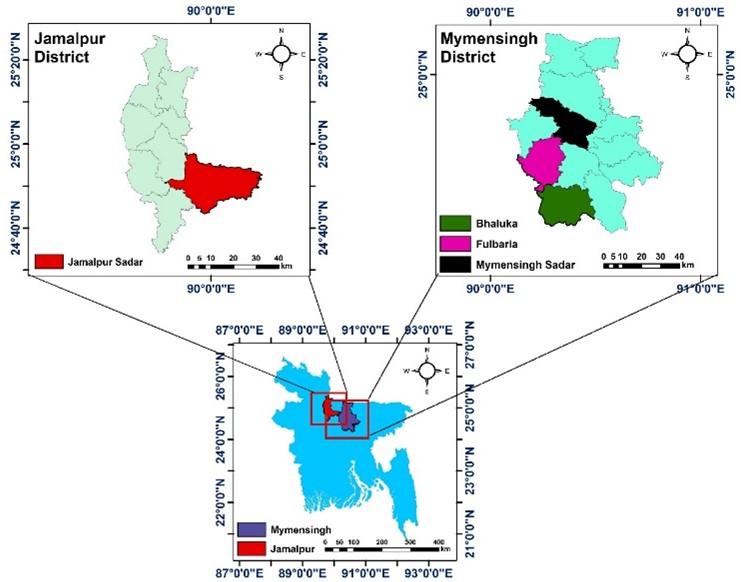
This study was performed from June 2018 to November 2019 in Mymensingh (24.7539° N, 90.4073° E) and Jamalpur (24.9250° N, 89.9463° E) districts of Bangladesh. The study areas are showed in Figure 1.
A total of 70 broiler samples comprising feces (n=20), chicken meat i.e. thigh, breast, and wings (n=30) from healthy birds, and visceral organs i.e. liver, lungs, and kidneys (n=20) from diseased birds were collected aseptically. Sterile cotton buds were used to collect freshly dropped fecal samples. Meat samples were collected by processing broilers from different markets. By post-mortem examination, visceral organs were collected from each bird that had lesions of avian salmonellosis. 5 gm of each samples was collected aseptically. Immediately after collection, samples were taken into sterile zip-lock bags with particular tag numbers and transferred to the laboratory maintaining a cool chain. After bringing to the laboratory, samples were seeded to sterile test tubes containing 5 ml sterile nutrient broth and incubated overnight at 37°C. All the experimental procedures and protocols used in this study were approved by the animal welfare and experimentation ethics committee of Bangladesh agricultural university (No. AWEEC/BAU/2019(28)).
Isolation of Salmonella spp.
Isolation of Salmonella spp. was performed by culture on Xylose Lysine Deoxycholate (XLD) agar (HiMedia, India) plates. Overnight enriched samples were streaked on XLD agar plates and incubated aerobically for 18-24 hours at 37°C to get pure colonies. Black-centered colonies on XLD agar plates were suspected as the growth of Salmonella spp. Gram’s staining and biochemical tests (urease test, sugar fermentation test, methyl red test, Voges-Proskauer test) were performed for further confirmation [18].
DNA extraction and PCR confirmation of Salmonella spp.
Isolated Salmonella spp. were finally confirmed by polymerase chain reaction (PCR) targeting the invA gene (F: 5′-ATCAGTACCAGTCGTCTTATCTTGAT-3′ and R: 5′-TCTGTTTACCGGGCATACCAT-3′) with 211 amplicon size [19]. For PCR, bacterial DNA was extracted by boiling and freeze-thawing method as previously described [20]. Briefly, initially 1 ml of overnight enriched culture was centrifuged at 5,000 rotation per minute (rpm) for 5 minutes and the supernatant was discarded. Subsequently, a similar process was performed after mixing 1 ml of phosphate buffer solution (PBS). After discarding supernatant, the pellet was suspended to 200 µL PBS; followed by boiling and cooling of the suspension for 10 minutes in each step. Finally, the suspension was again centrifuged for 10 minutes at 10,000 rpm and the supernatant was collected as genomic DNA. The collected genomic DNA was then stored at -20°C for further use.
A final volume of 20 µL consisting of 10 µL of the master mix (2X) (Promega, Madison, WI, USA), 4 µL of nuclease-free water, 1 µL of each primer, and 4 µL of genomic DNA (50 ng/ µL) was used to carry out the PCR amplification. The thermo-cycle conditions were as follows: initial denaturation at 95°C for 5 min, followed by 30 cycles of denaturation at 94°C for 30 s, annealing at 52°C for 2 min, extension at 72°C for 45 s, and final extension was conducted at 72°C for 45 s.
After amplification, PCR products were analyzed by 1.5% agarose (Invitrogen, USA) gel electrophoresis, stained with ethidium bromide (0.5 μg/ml) for 10 min in a dark place, and finally, the expected amplicon sizes were audited and captured under ultra-violet trans-illuminator (Biometra, Germany). A 100 bp DNA ladder (Promega, Madison, WI, USA) was used to check the targeted amplicon size.
Antibiotic susceptibility test
The antibiotic susceptibility test (AST) was done by the disk diffusion method [21]. Seven commonly used antibiotics under seven classes were employed: penicillins (amoxicillin- 30 μg), fluoroquinolones (ciprofloxacin- 5 μg), amphenicols (chloramphenicol- 30 μg), polypeptides (colistin- 10 μg), aminoglycosides (gentamicin- 10 μg), tetracyclines (tetracycline- 30 μg), and cephalosporins (ceftazidime- 30 μg). The AST was done by spreading freshly Salmonella growth culture having an equal concentration of 0.5 McFarland solution on Mueller-Hinton agar (HiMedia, India) plates. The guidelines of the clinical and laboratory standard institute [22] were followed to interpret the results. Any isolates showing resistance against three or more classes of antibiotics were deemed as MDR [23]. Furthermore, the multiple antibiotic resistance (MAR) index was evaluated by the following formula: MAR= a/b, where ‘‘a” denotes the number of antibiotics which were resistant to a particular isolate, and ‘‘b” denotes the total number of antibiotics tested [24].
Statistical analysis
Data obtained from this study were incorporated in Microsoft Excel-2010 (Los Angeles, CA, USA), and exported to the GraphPad Prism 8.4.2 (GraphPad Software, Inc.) and the Statistical Package for the Social Sciences (SPSS) software (IBM SPSS- version 25.0, USA) for statistical analysis. By SPSS, a Pearson chi-square test for goodness-of-fit was performed to observe the possible variations in the occurrence of Salmonella spp. and the resistance profiles of different antibiotics among different collected samples. Statistically significant p-value was less than 0.05. Furthermore, GraphPad Prism following the Wilson/Brown Hybrid method as previously described [25] was used to calculate the binomial 95% confidence intervals.
RESULTS
Occurrence of Salmonella isolates
Out of 70 samples, 46 (65.71%, 95% confidence interval: 54.04-75.75%) samples were positive for Salmonella spp. based on their colony characteristics and biochemical tests. Of these 46 isolates, 30% (21/70) samples were PCR positive for Salmonella spp. targeting invA gene; among which healthy broiler sample- feces (50%, 10/20) exhibited significantly higher occurrence of Salmonella spp., compared to internal organs (40; 7/20) from diseased broilers, and meat (10%, 3/30) samples from healthy broilers (Table 1).
Table 1. Occurrence of Salmonella spp. from different broiler samples.
Antibiogram profiles of isolates Salmonella spp.
From the antibiotic susceptibility test, all the Salmonella isolates were resistant to amoxicillin; frequently resistant to tetracycline (90.48%), ceftazidime (61.90%), chloramphenicol (38.10%), and colistin (33.33%). On contrary, gentamicin showed higher sensitivity to Salmonella isolates (Figure 2). Salmonella from visceral organs (diseased broiler samples) revealed peak resistance against most of the used antibiotics, where a statistically significant correlation was found for chloramphenicol, tetracycline, and ceftazidime (Table 2).
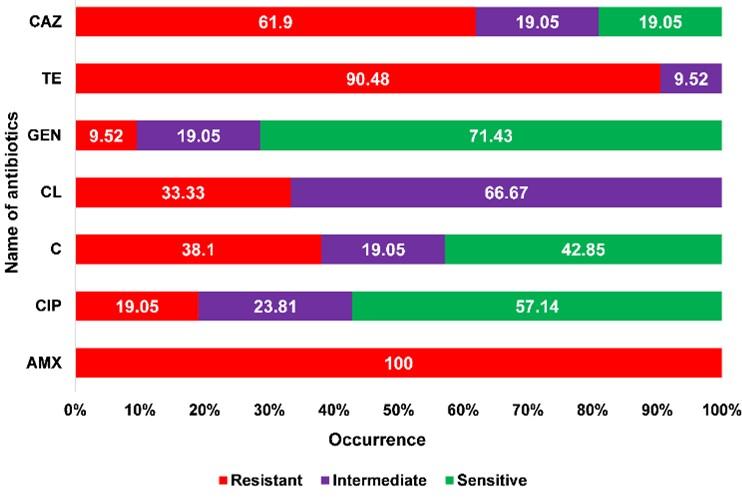
Table 2. Resistance profiles of Salmonella isolated from broiler samples.
Occurrence of MDR patterns and MAR index of Salmonella isolates
Out of 21 Salmonella isolates, 17 (80.95%; 95% CI: 59.99-92.33%) were MDR in nature. Overall, nine resistance patterns were audited, among them, the highest 23.53% (4/17; 95% CI: 9.56-47.26%) Salmonella isolates showed the resistance pattern no. 9 (AMX-TE-CAZ). One isolate showed resistance against six classes of antibiotics (six antibiotics) (pattern no. 1). The antibiotic resistance profile of each Salmonella isolate was found to vary with MAR indices ranging from 0.29 to 0.86. All the Salmonella isolates were resistant against at least two antibiotics representing two classes (Table 3).
Table 3. Occurrence of multidrug resistance and multiple antibiotic resistance index of Salmonella isolated from broiler samples.
DISCUSSION
Avian Salmonellosis is a major threat to both the poultry industry (causing serious economic losses) and human health (showing zoonotic significance). In addition, infections developed by MDR Salmonella spp. are difficult to control. Broiler meat, eggs, fecal materials, and visceral organs have been recorded as cardinal sources of Salmonella contamination [26]. Here, we reported the detection of MDR Salmonella from broiler chickens which show serious public health significance.
The invA gene of Salmonella usually comprises specific DNA sequences which proves the invA as a compatible gene to detect Salmonella genotypically [27]. In addition, the invA gene is available in almost all Salmonella serovars. This gene encodes a protein (inner membrane) that assists Salmonella to invade their epithelial cells [3]. In this study, the overall occurrence of Salmonella spp. targeting invA gene in broiler samples was 30% (21/70) which is lined with the previous study conducted in Bangladesh [15]. Conversely, both higher [5] and lower [16] prevalence rate of Salmonella spp. from broilers than our study were also recorded previously in Bangladesh. Globally, variable findings as 7.9% [26] and 0.75% [28] were recorded previously. This observed variations in the occurrence of Salmonella spp. might have linkage with the variations of the management systems of farms (biosecurity, hygiene, sanitary, etc.), sample size, types of samples, geographical and seasonal distributions, and method related factors. The occurrence of Salmonella in broilers suggests that the farms’ and poultry processing environments might contain poor-hygienic protocols. Furthermore, the presence of virulence gene invA in Salmonella isolates denotes their pathogenicity which can develop foodborne pathogens after introducing into food.
In the current study, a significantly higher occurrence of Salmonella spp. was observed in fecal samples (50%) of healthy broilers in relation to visceral organs (40%) of diseased broilers, and meat samples (10%) of healthy broilers. Previously several studies reported the presence of Salmonella spp. in broiler meat [29], fecal materials [5], and visceral organs [15]. The significantly higher occurrence of Salmonella in fecal materials is not unusual, as Salmonella are naturally found in the gastrointestinal tract of avian species [30]. These Salmonella contaminations can be introduced into the production system from broilers via feces, contaminated water or feed, and others. In addition, the presence of Salmonella in feces samples indicates that broiler droppings can shed Salmonella to other birds of the flocks. The presence of Salmonella spp. in meat samples denotes that Salmonella spp. have the potential to be transmitted to humans via the food supply chain. Furthermore, consumption of undercooked poultry and poultry products contaminated by Salmonella has also the potential in the transmission of Salmonella to humans [28].
Antimicrobial resistance is an emerging problem in the world and has the most significant public health challenge of this century globally [31]. Poultry and poultry products are huge sources of antibiotic reservoirs [32]. In our present study, all the Salmonella isolates were resistant to amoxicillin, and frequently resistant to tetracycline, ceftazidime, chloramphenicol, and colistin. Visceral organs exhibited a higher occurrence of antibiotic resistance compared to other selected samples (in the most antibiotics used). In addition, Salmonella resistance to tetracycline, ceftazidime, and chloramphenicol was significantly higher in visceral organs of diseased broilers. Interestingly, Salmonella isolates showed resistance to ceftazidime (61.90%) and colistin (33.33%) which is alarming for both human and animal health-care facilities. Ceftazidime is a 3rd generation cephalosporin antibiotic which usually used to treat severe bacterial infections in humans [33]. In addition, colistin is a reserved group of antibiotics which generally used only in severe infections developed by MDR Gram-negative bacteria [34]. However, MIC and molecular assays should be employed before drawing any conclusions.
Infections caused by MDR and MAR bacteria are serious global health concern as it is expensive for treatment and it may cause fatal consequences. MDR Salmonella has emerged as a cardinal human health issue throughout the world. The alarming situation was that 80.95% of Salmonella isolates were MDR in nature. Previously, Alam et al. [5] detected 100% MDR Salmonella spp. from broilers in Bangladesh. In addition, MAR indices of isolated Salmonella from our study were ranged from 0.29 to 0.86. More than 0.29 of MAR index denotes that antibiotics were frequently used in the sources from where Salmonella were isolated showing high-risk sources for MDR and MAR bacteria. The development of MDR and MAR in Salmonella may be the results of selective pressure triggered by the misuse and overuse of antibiotics in broilers [5]. These MDR and MAR Salmonella show severe public health significance by transmitting to humans through the food supply chain. In addition, these MDR and MAR bacteria can also spread in the environments and transfer their resistance genes to other bacteria horizontally.
CONCLUSION
High occurrence of MDR Salmonella spp. detected in our present study reveals a potential human and animal health risk. There is potential in the transmission of Salmonella spp. from broilers to one-health components through the food chain, and ultimately to contaminate them. Future studies including the detection of virulence and antibiotic resistance genes of Salmonella spp. from healthy and diseased broilers may clarify the actual dynamics of their transmission and dissemination to one-health components. Effective control strategies and sustained implementation of comprehensive risk reduction practices including strict biosecurity throughout the production continuum are required to minimize the emergence of MDR and MAR zoonotic Salmonella pathogens.
ACKNOWLEDGMENT
The authors are so grateful to farm owners for giving us access to samples during the whole study. The authors are also very much grateful to Dr. Khalada Zesmin, Upazila Livestock Officer, Kishoreganj, Bangladesh, for her valuable comments and suggestions during the whole study and the preparation of the manuscript. The authors are very much grateful to the Ministry of Education, Government of Bangladesh for providing funds through a research project (project number: LS2018686) to facilitate the present study.
AUTHORS CONTRIBUTIONS
Conceptualization, MFRK and MTR; Sample collection, MT and MSI; Methodology, MT and MSI; Software, MSI; Validation, MFRK and MTR; Formal analysis, MSI, MAS and MTR; Investigation, MT, MSI, SI and MN; Data curation, MSI and MT; Writing-original draft preparation, MSI and MT; Writing- review and editing, MSI, MAS, MFRK, FMB and MTR; Visualization, MSI, and MTR; Supervision, MFRK and MTR; Fund acquisition, MFRK and MTR; Critical revisions and writing, MFRK and MTR. All authors have read and agreed to the published version of the manuscript.
CONFLICTS OF INTEREST
The authors declare no conflict of interest.
References
- [1]Sabuj AA, Mahmud T, Barua N, Rahman MA, Islam MS, Bary MA. Passive surveillance of clinical poultry diseases in an Upazila Government Veterinary Hospital of Bangladesh. African Journal of Microbiology Research. 2019;13(29):632-639.
- [2]Islam MS, Sabuj AA, Haque ZF, Pondit A, Hossain MG, Saha S. Seroprevalence and risk factors of avian reovirus in backyard chickens in different areas of Mymensingh district in Bangladesh. Journal of Advanced Veterinary and Animal Research. 2020;7(3):546-553.
- [3]Tawyabur M, Islam M, Sobur M, Hossain M, Mahmud M, Paul S, Hossain MT, Ashour HM, Rahman M. Isolation and Characterization of Multidrug-Resistant Escherichia coli and Salmonella spp. from Healthy and Diseased Turkeys. Antibiotics. 2020;9(11):770.
- [4]Ievy S, Islam M, Sobur M, Talukder M, Rahman M, Khan MF, Rahman M. Molecular detection of avian pathogenic Escherichia coli (APEC) for the first time in layer farms in Bangladesh and their antibiotic resistance patterns. Microorganisms. 2020;8(7):1021.
- [5]Alam SB, Mahmud M, Akter R, Hasan M, Sobur A, Nazir KH, Noreddin A, Rahman T, El Zowalaty ME, Rahman M. Molecular detection of multidrug resistant Salmonella species isolated from broiler farm in Bangladesh. Pathogens. 2020;9(3):201.
- [6]Kirk MD, Pires SM, Black RE, Caipo M, Crump JA, Devleesschauwer B, Döpfer D, Fazil A, Fischer-Walker CL, Hald T, Hall AJ. World Health Organization estimates of the global and regional disease burden of 22 foodborne bacterial, protozoal, and viral diseases, 2010: a data synthesis. PLoS Medicine. 2015;12(12):e1001921.
- [7]Vindigni SM, Srijan A, Wongstitwilairoong B, Marcus R, Meek J, Riley PL, Mason C. Prevalence of foodborne microorganisms in retail foods in Thailand. Foodborne Pathogens and Disease. 2007;4(2):208-215.
- [8]Rahman M, Sobur M, Islam M, Ievy S, Hossain M, El Zowalaty ME, Rahman AM, Ashour HM. Zoonotic Diseases: Etiology, Impact, and Control. Microorganisms. 2020;8(9):1405.
- [9]Antunes P, Mourão J, Campos J, Peixe L. Salmonellosis: the role of poultry meat. Clinical Microbiology and Infection. 2016;22(2):110-121.
- [10]Aditya A. Drug resistant Salmonella in broiler chicken sold at local market in Bangladesh and its public health significance. African Journal of Biotechnology. 2015;14(43):2995-3000.
- [11]Islam M, Nayeem M, Hasan M, Sobur M, Ievy S, Rahman S, Kafi M, Ashour HM, Rahman M. Virulence Determinants and Multidrug Resistance of Escherichia coli Isolated from Migratory Birds. Antibiotics. 2021;10(2):190.
- [12]Davies J, Davies D. Origins and evolution of antibiotic resistance. Microbiology and molecular biology reviews. 2010;74(3):417-433.
- [13]Orubu ES, Zaman MH, Rahman MT, Wirtz VJ. Veterinary antimicrobial resistance containment in Bangladesh: Evaluating the national action plan and scoping the evidence on implementation. Journal of Global Antimicrobial Resistance. 2020;21:105-115.
- [14]Chereau F, Opatowski L, Tourdjman M, Vong S. Risk assessment for antibiotic resistance in South East Asia. Bmj. 2017;358:j3393.
- [15]Mridha D, Uddin MN, Alam B, Akhter AT, Islam SS, Islam MS, Khan MS, Kabir SL. Identification and characterization of Salmonella spp. from samples of broiler farms in selected districts of Bangladesh. Veterinary World. 2020;13(2):275.
- [16]Al Mamun MA, Kabir SL, Islam MM, Lubna M, Islam SS, Akhter AT, Hossain MM. Molecular identification and characterization of Salmonella species isolated from poultry value chains of Gazipur and Tangail districts of Bangladesh. African Journal of Microbiology Research. 2017;11(11):474-481.
- [17]Thrusfield, M. Veterinary Epidemiology, 2nd ed.; Blackwell Science: London, UK, 1995.
- [18]Sobur A, Hasan M, Haque E, Mridul AI, Noreddin A, El Zowalaty ME, Rahman T. Molecular detection and antibiotyping of multidrug-resistant Salmonella isolated from houseflies in a fish market. Pathogens. 2019;8(4):191.
- [19]Shanmugasundaram M, Radhika M, Murali HS, Batra HV. Detection of Salmonella enterica serovar Typhimurium by selective amplification of fli C, flj B, iro B, inv A, rfb J, STM2755, STM4497 genes by polymerase chain reaction in a monoplex and multiplex format. World Journal of Microbiology and Biotechnology. 2009;25(8):1385-1394.
- [20]Queipo-Ortuño MI, Colmenero JD, Macias M, Bravo MJ, Morata P. Preparation of bacterial DNA template by boiling and effect of immunoglobulin G as an inhibitor in real-time PCR for serum samples from patients with brucellosis. Clinical and Vaccine Immunology. 2008;15(2):293-296.
- [21]Bauer AT. Antibiotic susceptibility testing by a standardized single disc method. American Journal of Clinical Pathology. 1966;45:149-158.
- [22]Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing, M100-S28. Clinical and Laboratory Standards Institute, Wayne, PA. 2018.
- [23]Sweeney MT, Lubbers BV, Schwarz S, Watts JL. Applying definitions for multidrug resistance, extensive drug resistance and pandrug resistance to clinically significant livestock and companion animal bacterial pathogens. Journal of Antimicrobial Chemotherapy. 2018;73(6):1460-1463.
- [24]Krumperman PH. Multiple antibiotic resistance indexing of Escherichia coli to identify high-risk sources of fecal contamination of foods. Applied and environmental microbiology. 1983;46(1):165-170.
- [25]Brown LD, Cai TT, DasGupta A. Interval estimation for a binomial proportion. Statistical science. 2001;15(2):101-117.
- [26]Duc VM, Nakamoto Y, Fujiwara A, Toyofuku H, Obi T, Chuma T. Prevalence of Salmonella in broiler chickens in Kagoshima, Japan in 2009 to 2012 and the relationship between serovars changing and antimicrobial resistance. BMC Veterinary Research. 2019;15(1):1-8.
- [27]Yanestria SM, Rahmaniar RP, Wibisono FJ, Effendi MH. Detection of invA gene of Salmonella from milkfish (Chanos chanos) at Sidoarjo wet fish market, Indonesia, using polymerase chain reaction technique. Veterinary World. 2019;12(1):170-175.
- [28]Sedeik ME, Nahed A, Awad AM, Elfeky SM, Abd El-Hack ME, Hussein EO, Alowaimer AN, Swelum AA. Isolation, conventional and molecular characterization of Salmonella spp. from newly hatched broiler chicks. AMB Express. 2019;9(1):1-6.
- [29]Rizwan M, Yar SA, Achakzai SK, Shakoor M. PCR based detection of Salmonella from fresh and processed chicken meat from Quetta, Pakistan. International Journal of Biosciences. 2017;10(4):363-371.
- [30]Machado Junior PC, Chung C, Hagerman A. Modeling Salmonella Spread in Broiler Production: Identifying Determinants and Control Strategies. Frontiers in Veterinary Science. 2020;7:564.
- [31]Lambertini E, Ruzante JM, Chew R, Apodaca VL, Kowalcyk BB. The public health impact of different microbiological criteria approaches for Salmonella in chicken parts. Microbial Risk Analysis. 2019;12:44-59.
- [32]Hedman HD, Vasco KA, Zhang L. A Review of Antimicrobial Resistance in Poultry Farming within Low-Resource Settings. Animals. 2020;10(8):1264.
- [33]Gashe F, Mulisa E, Mekonnen M, Zeleke G. Antimicrobial Resistance Profile of Different Clinical Isolates against Third-Generation Cephalosporins. Journal of pharmaceutics. 2018;2018:5070742- 5070742.
- [34]Pogue JM, Ortwine JK, Kaye KS. Clinical considerations for optimal use of the polymyxins: a focus on agent selection and dosing. Clinical Microbiology and Infection. 2017;23(4):229-233.